►
From YouTube: Euler Cycles for “Life”
Description
Recorded presentation (10 minutes) to the TopoNets Satellite session of the Networks 2021 conference. Slides are available here: https://figshare.com/articles/presentation/Euler_Cycles_for_Life_developing_biological_structure_using_multi-cell_networks/14773362
A
Hello,
my
name
is
bradley
alicea,
there's
my
twitter
handle
and
I'm
from
the
open
room
foundation
and
the
divawarm
group.
Today,
I'm
going
to
be
talking
about
oiler
cycles
for
life,
so
this
talk
is
going
to
focus
on
euler
paths
and
euler
paths
are
ways
to
evaluate
a
network
by
looking
at
the
edges
instead
of
the
nodes,
and
so
euler
paths
are
sort
of
come
from
this
idea
of
the
seven
bridges
of
koningsberg,
which
is
shown
there,
and
you
can
apply
this
method
to
a
number
of
different
types
of
single
cell
multicellular
organisms.
A
So,
for
example,
you
have
diatom
cells,
which
are
single
cells
which
have
a
lot
of
internal
structure
or
volvox
colonies,
which
are
groups
of
cells
that
exist
in
a
colony,
and
so
we
can
take
these.
What
we
call
multi-cell
networks-
and
I
want
to
differentiate
between
the
multi-cell
idea
and
the
application
to
biological
cells.
Each
internal
region
of
a
network,
the
area
between
the
edges
is
considered
a
cell
and
biological
cells,
by
contrast,
can
be
analyzed
using
this
method,
so
you
can
have
multiple
network
cells
in
a
single
cell
or
a
network
cell.
A
That
combines
a
number
of
cells
within
it
or
it
can
represent
a
single
cell-
it's
pretty
flexible
in
that
way,
but
why
are
cell
boundaries
important?
So
if
we're
using
this
analogy
between
network
edges
and
cell
boundaries,
why
is
that
important
in
biology?
It's
very
important
for
a
number
of
reasons.
A
One
reason
is
because
it
serves
as
a
conduit
between
cells,
so
you
have
things
exchanged
like
mrna
molecules
or
ions
or
other
types
of
things
like
blood
flow
or
liquid
flow
or
oxygen.
Things
like
that
that
can
be
exchanged
within
in
between
cells
and
along
the
boundaries,
and
so
this
is
a
very
important
thing
to
understand
and
we'll
get
into
some
of
the
reasons
why
that
might
lead
to
multicellularity
later,
but
we're
going
to
evaluate
something
called
an
euler
circuit.
A
So
an
euler
circuit
is
a
set
of
edges
that
exist
in
a
network
and
it's
the
idea
that
we're
going
to
cross
every
independent
edge
of
us
in
a
multi-cell
network
once
and
only
once
so.
We
can
start
off
by
crossing
each
edge,
but
we
can
only
cross
each
edge
once
and
only
once
if
we
cross
an
edge
twice
or
if
we
failed
across
an
edge
within
our
network,
then
that
network
is
considered
to
be
an
incomplete,
euler
circuit.
A
A
So
let
me
show
you
how
you
evaluate
an
euler
path,
so
you
can
use
something
simple
like
this
graph
online
interface
and
you
can
build
a
network,
draw
it
out
and
then
find
the
shortest
path
in
that
network.
So
you
can
define
the
nodes
and
edges
and
then
define
the
path,
and
so
this
graph
has
an
eulerian
path,
that's
complete
and
it
shows
the
path
through
this
network.
A
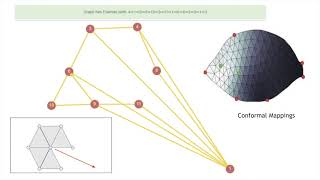
In
this
case,
though,
this
network
does
not
have
a
complete
euler
path,
and
so
this
is
shown
in
the
interface
as
being
an
incomplete
path.
It
doesn't
tell
you
how
many,
how
many
steps
it
missed
or
how
many
steps
it
exceeded
it
by,
but
we'll
just
say
that
it
doesn't
have
an
eulerian
solution,
and
so
this
this
shape.
I
want
to
take
this
and
actually
get
to
another
issue,
which
is
that
we
can
take
one
of
these
networks
and
deform
them.
A
A
So
a
number
of
tests
that
we
can
do
to
show
how
this
technique
works
in
terms
of
evolving
phenotypes
and
evolving
multicellular
systems
comes
from
testing
out.
Our
triangle
motif
so
in
this
case
we're
going
to
build
a
lineage
tray
of
these
multi
cells,
we're
going
to
start
with
a
triangle
and
then
we're
going
to
add
another
triangle
on
on
the
side.
So
we're
going
to
have
a
simple
replication
rule,
but
we
find
that
that
simple
replication
yields
an
incomplete
oil
or
path.
So
now
we
have
a
choice.
A
We
can
either
add
in
a
secondary
shape,
which
is
this
hexagon
and
that
actually
restores
our
original
euler
path,
but
it
introduces
something
called
modularity,
which
means
that
now
it
has
a
separate,
it
has
a
new
sort
of
piece
to
it.
That's
going
to
replicate
and
it's
going
to
be
a
differentiated
morphology
or
we
can
add
another
triangle
and
we
can
still
end
up
with
an
incomplete
euler
path
and
then,
of
course,
we
can
add
additional.
A
We
can
have
a
simple
replication
rule
which
gets
us
further
away
from
the
oil
complete
euler
path,
which
we
can.
Then
you
know
remedy
by
introducing
these
new
shapes
in
the
middle
and
having
this
new
modularity
that
emerges,
and
you
can
do
the
same
thing
here
in
this
path,
where
you
add
a
shape
in
a
different
way.
A
So
then
you
have
modularity
as
well.
So
basically
what
happens
is
when
you
have
an
incomplete
euler
path
in
these
lineage
trees.
You
add
something
to
see
what
recovers
the
euler
completeness
and
then
that
usually
leads
to
something
called
modularity
or
to
have
like
a
differentiated
phenotype
with
two
or
three
different
modules
in
it,
and
so
this
is
kind
of
what
we
see
with
diatoms,
where
they
have
these
triangular
shapes
or
other
shapes,
and
then
they
grow.
They
accrete
their
body
and
you
know
they're.
A
And
so
then
we
can
do
the
same
thing,
though,
with
a
rectangular
shape
and
a
simple
replication
rule
in
our
transformation
rule.
So
now
in
this
tree
we
have
a
simple
rectangle.
We
have
a
replication
rule
which
preserves
that
euler
completeness.
We
have
another
duplication
rule
which
preserves
that
oil
or
completeness
but
an
asymmetric
transformation.
Even
that
actually
keeps
that
oiler
completeness.
A
So
this
is
actually
a
pretty
robust
shape,
but
then,
finally,
we
get
to
a
point
where
we
do
this
h
asymmetric
transformation
twice
and
we
end
up
with
something:
that's
not
oil
or
complete.
So
this
lineage
is
not
going
to
be
oil
or
complete.
However,
this
one
here
you
keep
going
and
you
add
in
you
know,
maybe
you
add
in
another
a
new
shape
and
you
end
up
losing
your
oil
or
modular
or
your
oil
or
completeness,
and
then
eventually
you
get
to
this
modularity,
where
you
recover
it.
A
In
this
case,
we
have
in
this
branch
of
the
tree.
We
have
fractal
reproduction.
So
this
is
the
case
I
talked
about
earlier,
where
we
have
replication
rules
that
occur
recursively
and
we
have
this
fractal
structure.
So
it's
it's,
it
emerges
at
different.
You
know
numbers
of
cells,
it
always
retains
that
euler
completeness
and
you
have
things
that
have
maybe
very
little
internal
structure,
but
you
know
grow
to
be
very
large.
A
This
motif,
and
so
again,
hexagons
are
very
good
at
this,
and
so
this
might
explain
some
of
the
things
in
nature
where
you
have
cells
that
just
keep
replicating
and
they
don't
somehow
form
like
things
that
are
hugely
complex.
They
just
kind
of
grow
and
grow
and
grow,
and
so
that
you
know
there
are
ways
we
want
to
continue
with
this
type
of
work.
To
see
what
kinds
of
shapes
we
can
find,
what
kind
of
roles
we
can
find
and
so
forth.
So
I
wanted
to
put
a
plug
in
from
growth
and
form.
A
So
this
is
the
direction
we
want
to
go
in
to
sort
of
look
at
on
growth
and
form
in
a
new
way
and
put
numbers
on
it
and
think
about
it
in
terms
of
network
theory,
thank
you
for
your
attention
and
join
us
in
our
groups,
our
dvoram
group
and
our
openworm
foundation.
You
can
join
our
slack
with
that
link
or
you
can
visit
us
on
the
web
at
that
link.
