►
From YouTube: DevoLearn Tutorial (13 minutes)
Description
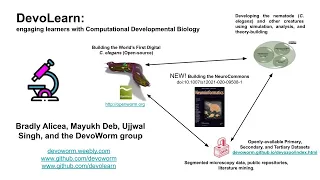
A tutorial on the DevoLearn platform, April 2021. Presented at the INCF Assembly (virtual, 2021). Contributors: Bradly Alicea, Mayukh Deb, Ujjwal Singh, and DevoWorm group, especially our Google Summer of Code students and applicants.
A
This
is
a
project
that
I've
worked
on
along
with
mayuk
deb
lujwal
singh
in
the
diva
worm
group
mayo
can
usual
were
jisock
students
in
2020,
and
so
this
project
grew
out
of
their
efforts,
but
also
the
efforts
of
previous
gsoc
students.
In
previous
years.
You
can
visit
our
website
at
divaworm.weebly.com
and
there
you'll
find
a
list
of
our
other
collaborators,
and
people
contributed
in
different
ways.
A
A
A
We
use
secondary
data.
We
use
other
types
of
approaches
to
build
simulations,
analog
data,
analysis
and
theory
building
to
understand
development.
Better
often,
we
can
talk
about
ourselves
as
a
computational
developmental
biology
group
or
data
science
biology
group.
Finally,
we
are
very
invested
in
open
data,
our
entire.
A
We
have
some
collaborators,
but
a
lot
of
our
activities
are
based
around
open
data,
and
so
we
have
this
platform
devo
zoo,
where
you
can
find
openly
available.
Primary
secondary
tertiary
data
sets
all
annotated
and
put
together
in
a
way
for
people
to
come
to
the
group
and
use
so
this
includes
data
such
as
segmented,
microscopy
images,
public
repositories
of
different
types
of
data
and
literature
mining.
Although
we
have
a
broader
array
of
data
that
are
available
to
us.
A
A
So
this
is
the
diva
learn
platform
and
you'll
find
it
at
github.com
divalern,
and
it
consists
of
three
parts.
So
the
first
part
is
this:
diva
learn
0.3.0,
and
this
is
the
standalone
diva
learn
program.
It's
a
pre-trained
model
for
machine
learning
for
deep
learning,
so
it
takes
data
and
it
does
a
sort
of
it
pre-trains
the
model
so
that
you
don't
have
to
train
it
with
labels.
A
A
A
A
One
of
the
more
exciting
projects
that
we
have
going
is
the
sort
of
a
neural
cellular
automata
model,
and
so
this
is
where
you
combine
neural
networks
or
deep
learning,
networks
and
cellular
automata
and
of
course,
cellular
automata
are
a
good
way
to
model
morphogenesis,
and
so
this
is
yet
another
thing
that
we're
going
to
be
putting
into
this
platform.
So
we're
constantly
building
up
this
platform,
you
can
find
divorm
ai
at
evwarm.github.io
divormai.
A
So
then,
the
final
part
of
the
stevenlearn
platform
is
the
devozu,
and
so
the
devozu
is
this
collection
of
secondary
data.
It's
organized
by
species,
and
we
give
you
publicly
available
data
sets,
we
annotate
everything
and
we
give
you
a
list
of
things
to
download,
and
so
anyone
can
come
into
our
group
and
conduct
an
analysis
and
it
might
be
a
comparative
analysis.
It
could
be
an
analysis
of
a
certain
species
and
you
know
they're
whatever
they
would
like
to
do.
A
So
we
have
this
further
goal
of
looking
at
quantitative
morphology,
using
deep
learning,
and
we
we're
doing
this
in
a
number
of
ways,
as
I
said
before,
there's
the
devo
zoo,
but
we
also
have
these
different
target
models
that
we're
going
to
be
looking
at
it
now
and
in
the
next
couple
years.
The
first
is
a
model
of
axolotl
brain
development,
so
we
have
a
collaborator
who's.
Collecting
data
on
axolotl
embryos
and
the
thing
about
axolotl
embryos
is
that
they
have
a
transparent
surface.
A
So
you
can
see
the
brain
as
it
develops
in
these
embryos,
and
so
this
is
the
embryo
at
the
top,
and
this
is
the
acts
the
fully
grown
axolotl
at
the
bottom,
and
you
can
actually
observe
this
process
and
our
collaborator
is
able
to
acquire
these
images
of
axolotl
embryos
using
a
number
of
really
innovative
techniques.
For
example,
she
has
a
ball
microscope
which
takes
images
from
different
angles
of
the
surface.
A
She
also
has
a
microscope
that
inverts
the
embryo
using
a
water
column,
and
so
all
these
things
are
available
for
people
to
analyze.
Further,
you
can
see
the
c
elegans
embryogenesis
example
on
the
bottom.
This
is
a
screenshot
from
the
devo
learn
0.3.0
software.
This
is
where
you
have
you
have
you
can
build
these
animations?
A
You
can
segment
the
cells
in
the
build
animation,
so
you
get
these
time
lapse
images
or
these
time
series
of
an
embryo,
or
you
know,
looking
at
different
frames
of
the
acquisition
of
the
embryo,
and
so
you
and
you
can
also
put
it
into
a
coordinate
system,
much
as
like
you
might
do
with
cell
tracking.
So
you
can
reconstitute
cell
tracking
in
this
way,
you
can
also
look
at
diatoms
and
the
species
of
dye
or
the
genus
that
diatom
we're
interested
in
is
basil
area.
A
A
We've
developed
a
technique
to
pull
apart
some
of
these
images
and
look
at
how
they
sort
of
move
relative
to
one
another
in
the
colony
and
we're
getting
these
images
from
another
collaborator.
Who
is
able
to
do
some
actually
pretty
interesting
microscopy
with
respect
to
this
organism?
So
our
sort
of
our
end
goal
is
to
conduct
theory
building
or
maybe,
like
you
know,
straightforward
analyses,
testing,
hypotheses
to
explain
developmental
processes,
and
you
can
see
we're
interested
in
this
process
where
you
go
from
a
sphere
to
this
differentiated
organism.
A
So
I
mentioned
before
that.
This
is
google
summer
code
is
an
integral
part
of
this
platform,
developing
it.
So
we've
been
preparing
for
google
summer
of
code.
This
is
where
we
provide
education
and
evaluation
for
students
in
deep
learning
and
machine
learning,
but
also
people
who
are
interested
in
developmental
biology
may
not
have
the
background
to
join
a
lab,
and
so
we
offer
this
sort
of
bridge,
and
so
by
you
know,
getting
involved
in
the
diva
owner
platform
is
an
excellent
way
to
do
that.
A
We've
made
calls
for
involvement
and
some
of
those
have
involved
g
suck
and
some
of
those
have
involved
just
regular
calls
for
involvement.
People
will
contribute
to
a
github
issue
during
the
application
period
or
whenever
they
want.
We've
had
a
lot
of
contributors
so
far,
this
gsox
season
where
people
are
trying
to
apply
for
their
projects
and
in
the
process,
they
issue
a
number
of
pull
requests,
and
my
oak
has
been
good
enough
to
keep
on
top
of
that
and
maintain
these
pull
requests.
A
And,
finally,
they
get
to
join
our
weekly
meetings.
We
have
a
meeting
once
a
week
and
this
helps
them
to
get
involved
in
bigger
projects
to
gain
a
better
perspective
on
what
they're
doing
in
the
code
base.
So
if
they're
just
issuing
a
pull
request,
you
know
you
can
see
the
code
and
you
can
see
the
net
the
deep
learning
problem,
but
you
don't
actually
know
why
and
so
we're
sort
of
bridging
that
gap
for
students
and
so
this
divalerm
package.
A
This
is
in
an
area
we
call
sort
of
data
science,
or
you
know
some
sort
of
collection
of
demos
that
people
would
need
if
they
want
to
say,
for
example,
take
the
next
step
from
the
deep
learning
part
which
is
like
you
know,
segmenting
images
and
finding
pattern
broad
patterns
in
the
data
to
something
like
doing
a
data
analysis,
and
they
can
learn
that
in
this
platform
we
also
have
educational
resources.
So
we
look
at
teaching
computational
students
about
developmental
biology,
so
we
have
in
some
of
our
devo
zoo
materials.
