►
From YouTube: 6. KBase and Jupyter
Description
June 11, 2019 Jupyter Community Workshop talk by Bill Riehl, Lawrence Berkeley National Laboratory
A
My
name
is
bill:
real
I
work
here
at
Berkeley,
National,
Lab
and
I'm
gonna
talk
about
the
do
a
systems
biology
knowledgebase,
which
we
use
the
jupiter
notebook
as
one
of
the
main
front
ends
for
for
users
to
come
in
and
do
their
analytical
work.
So
at
first
what
is
cave
a
soak
a
base
is
a
knowledge
creation
and
discovery
environment,
so
I'm
for
both
biologist
and
bioinformatics
ist's.
So
the
more
comprehensive
version
of
what
that
means
is
any
any
vial
that
you're
any
biologists.
A
Anybody
who
has
biological
data
can
come
in
into
the
cave.
A
system
upload
their
data,
use
our
resources
to
do
a
number
of
different
balances,
pipelines
on
that
and
check
out.
The
results
interpret
those
results
document
that
interpretation
and
share,
including
the
the
notebook
that
they
would
use
to
do.
A
As
focused
around
environmental
interactions,
so
environmental
biology,
microbial
biology,
plant
biology
and
Ike
Rovio
communities,
so
users
can
come
with
that
from
GE
user
facilities
or
even
things
that
they
get
garnered
from
NCBI
up
front
uploaded
into
our
system,
and
that's
not
just
as
simple
as
throwing
data
files
that
they
have
on
their
laptops
into
a
web
page.
So
what
we
mean
by
data
integration
at
this
step
is
kbase
is
built
around
a
pretty
strong,
centralized
data
model.
A
A
Is
we
really
want
to
build
this
as
a
knowledge
base,
so
a
way
to
react,
data
of
different
types,
and
that
comes
from
different
experiments
and
from
analyses
we
integrate
all
of
that
data
together
and
try
to
build
knowledge
and
try
to
be
able
to
build
predictive
biology
out
of
all
that,
it's
pretty
lofty
goal,
and
after
after
some
time
of
getting
getting
the
basics
down,
we're
we're
finally
making
headway
towards
that.
But
I
won't
have
much
time
to
get
into
that
today.
A
We
can
chat
in
the
breaks
if
you
like
anyway,
once
data
gets
uploaded
and
integrative.
The
next
step
is
to
do
this
collaborative
analytics.
We
call
where
users
can
come
in
use,
apps
and
tools
that
we
provide
or
even
provide
apps
and
provide
their
own
tooling
to
kabe
a
system
to
do
analysis
of
data,
and
the
way
that
this
is
done
is
through
a
fairly
heavily
modified
version
of
the
Jupiter
notebook.
A
A
All
of
this
again,
all
of
this
together
I
just
want
to
reiterate
the
the
data
that
a
user's
uploaded
and
transform
to
me
to
the
data
model,
as
well
as
the
notebook
itself
and
the
analysis
itself,
and
even
the
job
status
and
job
documents
themselves
get
bundled
together
and
become
themselves
a
shareable
unit
in
kbase.
So
if
I
upload
things
and
work
on
them,
I
can
share
them
with
Shane
or
with
Rowling
or
anybody
here
who
can
also,
at
the
same
time
perform
different
analyses.
A
Take
notes
in
that
and
I'll
be
alerted
to
see
when
those
are
updated
and
we
are.
We've
also
recently
introduced
a
way
to
build
up
user
groups
and
the
more
broader,
not
just
within
the
notebook,
but
throughout
the
whole
system,
way
to
manage
data
and
work
slowly.
Judging
for
zhen
shan,
mr
before
and
I
know,
there's
other
people
struggling
with
the
concept
or
struggling
with
the
practice
of
being
a
truly
fair
data
share
organization.
A
As
mentioned,
we
have
a
series
of
apps
that
can
be
run
on
a
gambit
of
biological
data.
We
have
about
200
right
now
and
it's
it's
very
much.
Not
a
closed
system
kbase
is
a
very
open
platform,
so
anybody
who
has
and
any
app
and
the
external
app
or
anything
that
they
want
to
be
able
to
use
on
our
resource
or
integrate
with
our
data
model.
There's
an
open,
SDK
and
that's
available
to
use.
It's
really
made
for
the
community
a
little
bit
about
the
the
architecture
behind
kbase.
A
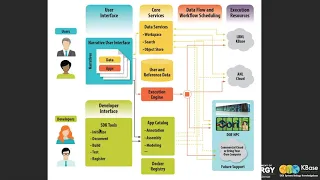
So
our
interface,
which
we
call
the
narrative
interface,
it
is
built
on
the
Jupiter
notebook
good
binds
to
the
data
and
the
apps
and
the
analysis
together
behind
all
that
core
services.
The
main
data
service
that
might
be
of
interest
is
sitting
on
MongoDB
and
that,
on
top
of
itself,
on
top
of
Apple
App
Store,
where
the
larger
chunks
of
data
like
meta-genome
reads
installing
are
stored
as
well
as
user
and
reference
data.
A
Next
bit
is
the
the
execution
engine,
so
the
once
a
cell
is
clicked
or
a
cell
is
run
that
contains
an
app
in
it
that
gets
fired
off
and
sent
to
the
execution
engine
which
runs
asynchronously
and
alerts
the
notebook.
When
it's
finished
so
then
the
notebook
will
update
itself
from
developer
interface.
I
also
won't
touch
on
too
much,
but
we
do
have
a
set
of
SDK
tools
for
adding
your
own
apps
to
the
system
and
just
it's
pretty
open
and
free
to
use.
A
So
you
could
just
plug
it
right
into
the
app
catalog
and
then
it'll
become
available
for
anybody
who
would
want
to
use
your
app
and
I'll
do
a
brief
demo
here.
I
think
some
of
the
concepts
make
more
sense
to
really
see
it
in
action.
So
I
have
one
up
right
now.
This
is
the
tutorial
I
have
on
the
jdi's
meadow
genome
assembly
pipeline
that
we
have
wrapped
up
as
a
single
app
in
cádiz
and
there
this
is
a
Jupiter
notebook.
It's
pretty
heavily
modified.
A
We
have
a
different
set
of
templates
that
work
on
the
front
end
that
give
us
things
like
this
data
panel
here.
So
this
is
the
set
of
data
objects
that
are
associated
with
this.
This
narrative
there's
an
each
one
of
these
is
itself
tell
you
what
it's
data
type.
Is
this
one's
an
RNA
seek
alignment
a
paired
end
library
reads:
opening
it
up
will
give
you
some
options
and
tell
you
quite
a
bit
about
the
metadata.
A
You
can
also
drag
and
drop
on
here,
which
I
want
you
right
now,
so
I
don't
want
to
eat
up
the
network,
but
that
will
also
that
will
just
automatically
create
itself
actually
yeah
I'll
gamble,
let's
gamble
so
create
this.
Will
pop
a
cell
in
place
that
once
this
loads
up,
it
will
show
you
a
little
bit
about
what
that
data
object,
is
and
give
you
some
details
and
I
just
want
to
emphasize
a
little
bit
we're
not
making
up
any
new
cell
types
here.
These
are.
This
is
a
standard
Jupiter
notebook.
A
A
Scrolling
down
a
little
bit
when
the
user
gets
to
the
point
that
they
would
want
to
run
an
app
there's,
what
we
call
an
app
cell,
which
again
is
just
another
code
cell,
and
this
just
gives
a
different
interface
really
for
a
user
to
create
code.
So
if
there's
power
user-
like
probably
everybody
in
this
room,
you
can
just
enter
code
directly
and
execute
it,
but
a
number
of
our
external
users
that
are
pure
biologists
or
pure
bench
viola
districts.
A
Para
mental
biologists,
aren't
necessarily
interested
in
writing
the
code
themselves,
but
so
we
also
provide
an
interface
that
this
cell
becomes
very
well
aware
of
what
data
is
available
in
this
narrative.
So
you
can
kind
of
pick
and
choose
what
what
I
want
to
run
on
for
my
inputs
decide
what
the
output
should
be
and
then
just
hit
run,
that'll
execute
and
as
the
couldn't
show
style.
This
would
be
the
result
of
that.
So
there's
two
other
things
that
become
active
here.
A
A
Finally,
so
what's
next
for
us,
is
that
was
all
living
on
the
Jupiter
notebook
right
now
and
actually
a
little
bit
of
an
older
version
of
that
as
well,
but
we're
all
pretty
excited
about
Jupiter
lab,
especially
coming
out
1.0
very
soon.
Congratulations
guys!
So
one
project
that
we
have
in
mind
is
we
want
to
adapt
what
are
currently
a
series
of
env
extensions
into
the
series
of
droop.
Your
lab
extensions
we've
started
that
work,
and
some
of
it
will
be
challenging,
so
I
might
be
bugging
some
of
the
Jupiter
folks
that
are
here
today.
A
We're
also
using
we're
also
transitioning
over
to
using
Jupiter
hub
for
detainer
management
for
notebooks
for
various
other
things
yeah.
We
have.
We
have
a
custom
system
right
now
that
that
spawns
notebooks,
if
every
notebook
or
narrative,
probably
stick
with
the
byline
narrative
that
you
see
in
the
narrative
interface
is
a
docker
container,
that's
running
in
for
each
individual
user
and
we'll
be
transitioning
that
over
to
be
using
a
tooth,
you
want
to
hub
keep
tripping
over
those.
A
Partly
I
think
it
will
be
well
there-there's.
A
few
things
we've
talked
about,
so
one
is
one
of
the
great
things
about
Jupiter.
Lab
I
think
is
that
everything
is
built
in
as
an
extension,
so
we
can
really
build
a
flavor
of
Jupiter
lab
that
will
look
more
or
less
like
what
the
narrative
looks
like
now.
Every
I
mean
we
already
have
some
progress
on
I
think
I.
Have
that
open
my
little
act
together
version,
so
this
is.
A
We
already
have
an
extension
that
has
here's
the
list
of
apps
that
we
have
here's
the
set
of
data
that's
available
in
a
given
notebook,
notebooks
aren't
really
grey
skin.
Yet
you
have
the
the
file
browser
links
to
our
data
store
in
Albany,
so
it's
already
kind
of
on
the
way
towards
looking
like
that.
What
is
different.
A
Your
right
is
a
lot
of
the
other
branding
components,
but
my
feeling
is
that
Jupiter
labs
sensible
enough
that
that
will
be
quite
a
bit
of
work,
but
I
think
mostly
will
be
doable
in
the
world
also.
This
is
also
an
opportunity
for
us
to
update
what
our
look
and
feel
should
be,
and
we've
learned
a
lot
of
lessons
the
past
few
years
of
what
works.
What
really
doesn't.
