►
From YouTube: Open Source Antibiotics Science Update Apr 6th 2022
Description
Weekly open project meeting for Open Source Antibiotics Series 2.
Full Project: https://github.com/opensourceantibiotics/Series-2-Diarylimidazoles
Relevant GitHub Issue: https://github.com/opensourceantibiotics/Series-2-Diarylimidazoles/issues/114
On the call: Professor Matthew Todd, Dr Edwin Tse, Dr Daniel Gedder (UCL), Prof Lee Graves, Thomas Gilbert (UNC), Dr Lori Ferrins, Maria Dichiara (NEU), Dr Chris Swain (Cambridge Medchem Consulting).
A
B
Okay,
so
today's
meeting
wednesday
6th
of
april
the
I
guess,
the
two
things
we
were
hoping
to
talk
about
were
data
from
lee's
lab
and
then
compounds
from
daniel
and
potentially
from
maria,
so
thinking
about
what
we
might
be
testing
next
and
potentially
last
before
we're
trying
to
write
up
the
paper.
So
those
are
the
two
things
today
really
and
so
lee
tom
did.
You
want
to
dig
right
into.
A
Yeah
I've
I've
prepared
some
slides
just
to
go
over
really
quickly.
You
know
I,
as
you'll
see
this
has
taken
a
lot
longer
than
we
wanted,
of
course,
with
coven
and
everything,
but
I
think
we're
turning
the
corner.
I
feel
like
there's
a
few
things
we'd
like
to
repeat
and
I'll
talk
about
that,
but
I
feel
like
maybe
we're
getting
some
insight
and
I'm
hopeful
anyway.
A
A
Everybody
see
that
okay,
I
hope
that's
good
right
and
the
idea
that
we're
adapting
our
approach
to
is
the
competition
assay,
where
basically,
we
incubate
with
the
compounds
and
then
ask
if
we
compete
binding
to
the
inhibitor
beads
and
look
at
subtractive
analysis.
Basically,
if
we've
competed
off
something
and
we
try
to
do
it
in
a
dose-dependent
manner-
and
we've
used
this
quite
extensively
to
you
know
to
determine
kinase
inhibitor
selectivity
in
the
past.
A
So
if
you
remember
the
first
attempt,
we
got
a
mrsa
lysate,
we
weren't
very
happy
with
the
results
it
seemed
like
we
were
struggling
with
enough
mercy
lysate.
So
we
were
also
concerned
if
the
assay
was
working
properly.
So
what
what
we
did?
I
should
just
reiterate:
the
goals
of
the
project
were
to
profile
compounds,
26
and
28
in
mrsa
and
then
also
to
profile,
26
and
28
against
the
human
kinome
and,
as
I
said,
because
we
were
struggling
with
our
initial
analysis
of
the
mrsa
data.
We
weren't
sure
that
everything
was
working.
A
B
A
The
point
is
that
you
know
we
have
an
assay
that
works.
We
have
some
insight
and
I
and,
as
I
said
before,
I'd
like
to
repeat
this
experiment,
so
we
have
at
least
two
independent
verifications
of
this,
because
I
think
it
could
be
important
in
terms
of
what
the
specificity
is
and
what
the
potential
effect
on
human
cells
is
now
most
recently,
you
know,
as
many
of
you
know,
the
bottleneck
has
been
getting
safe
and
fortunately
we
found
a
lab
next
door.
A
A
Which
is
an
atp
binder,
and
so
none
of
those
three
proteins
changed
significantly
with
26,
so
they
were
very
much
28
dependent.
There
were
other
ones
that
were
26
dependent,
which
I
think
we're
not
so
interested
in
those
as
26.
I
think
it's
the
control
compound,
but
we
were
interested,
and
I
did
a
little
bit
of
research
on
these
three
and
again,
there
were
others
that
we
could
potentially
tease
out
of
there
that
were
much
weaker.
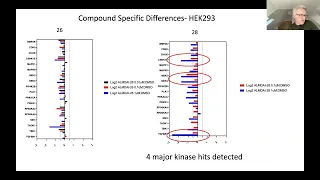
A
These
were
among
the
strongest
hips
that
were
found
and
and
so
results
I
would
say,
of
the
kinases
we
can.
We
can
breathe
pretty
certain
that
we're
seeing
competition
against
probably
casein
kinase,
one,
the
two
neck
kinase
and
the
tgfr
beta
kinase
that
I
mentioned,
and
then
in
the
immersive
proteome.
A
A
If
it's
inhibited
could
potentially
be
important
in
terms
of
regulating
protein
translation
in
mrsa
and
then
this
mnma
is
a
trna,
specific
thiourdylic
delays
and,
like
I
said
it's
an
atp
binder,
and
there
was
a
paper
from
a
group
of
acs
recently
that
talked
about
this
potentially
being
down
regulated
as
a
result
of
lactobionic
acid
treatment
which
is
being
tested
for
mercy.
So
all
three
of
them
have
some
credibility
to
them
in
terms
of
potential
potential.
Targeting,
I
think
what,
like
I
said,
what
I'd
like
to
do
is
then
figure
out.
A
A
Are
future
directions
yeah?
So
I
think
the
first
thing
we
should
do
is
repeat
our
hdk
293
kind
of
analysis
just
to
get
better.
You
know
values
on
everything
again,
so
we
can
say
for
certain
that
we
think
these
kinases
are
likely
to
be
targeted
by
the
28
and
not
26.,
I'd
like
to
repeat
the
mercy
experiment,
one
more
time
we
can
get
more
mercilessate
from
brian
conland's
group
and
just
go
ahead
and
repeat
that
and
see.
A
So
many
thanks
to
kareem
he's
really
carried
this
project
all
the
way
through
thanks
to
brian
conlon's
lab
for
stepping
in
and
getting
us
mrsa
lysate,
which
is
not
easy
to
do.
And
then
laura
herring
and
the
proteomics
group
for
running
the
analysis
on
this
and
happy
to
take
any
questions
or
comments.
B
All
right,
thank
you,
that's
fantastic,
so
yeah
the
there
were
the
two
repeats
there
that
you're
talking
about
are
they
I
mean
I
mean
given
that
they
repeat.
So,
are
you
able
to
do
those
in
the
reasonably
short
term?
I
mean
how's
your
pipeline,
looking
of
experiments
that
you
ought
to
do
well,
we're
extremely
busy.
A
A
But
I'll
just
have
to
check
and
see
what
the
time
we
have
tons
of
293
cell
lysates
in
the
freezer,
which
we
can
break
open
and
do
quite
quickly
and
cream,
might
even
be
able
to
get
our
new
person
to
do
those
experiments.
But
I
actually
probably
have
him:
do
it
just
because
he's
more
experienced
but
yeah
that's
what
cream.
B
C
B
C
B
A
Well,
we
could,
and
but
in
some
ways,
I'd
like
to
just
go
ahead
and
do
the
whole
thing
again
just
so
that
we'd
have
two
completely
independent
experiments
to
say
you
know
this
is
what
we
found
and
we
could
certainly,
then
I
I'm
sure
david
brewery
has
neck
and
antibodies.
We
could
do
some
simple
experiments
to
see
you
know
just
competition
binding
or
something
with
an
antibody
to
validate
that
it
would
make
a
nice
figure,
the
tgf
beta
r1.
A
I
don't
know
what
you
think
about
that
and
I,
I
suspect
we
should
pay
attention
to
that
as
well,
since
that
was
a
pretty
strong
hit.
In
fact
that
was
the
strongest
hit
we
had.
It
was
clearly
different
from
28
26.
I
mean
26
didn't
touch
it
right.
So
in
terms
of
biology,
I
don't
know
what
to
make
of
that
either,
but
and
and
maybe
lori
laurie.
Can
you
tell
us
more
where
what's
the
origin
of
these
compounds
over
these
things
that
you
made
has
any
other
profiling
been.
E
B
A
Well,
I
know
they
have
a
great
interest
in
neck
kinase,
and
so
it
wouldn't
surprise
me
if
they
have
some
nick
effect.
There
are
other
kinases
in
here
that
did
show
some
response.
That
might
be
important.
Plk
pka
there's
a
little
bit
of
a
response,
and
I
think
if
we
just
repeat
the
experiment
again,
we
can
say
with
greater
confidence.
We
can
potentially
expand
this
list
a
little
bit
more
than
an
n
of
one.
B
B
A
B
B
A
A
A
B
A
B
A
Right
and
the
mnma
is
clearly
an
atp
binder,
so
it
would
make
sense
why
it
might
bind
to
these
beats
the
other
two.
It
wasn't
clear
to
me
that
there
was
a
atp
or
nucleotide
pocket
that
might
determine,
although
possibly
there
is
something
that
you
know
would
allow
them
to
bind.
You
know
in
a
bead
specific
manner
that
could
show
competition.
A
Yeah
in
some
ways,
I'd
be
very
pleased
if
we
find
some
new
targets
to
this,
because
if
it's
a
challenge
right
with
these
are
designed
for
kinases,
if
we
capture
things
that
are
not
kinases
and
we
can
show
competition,
ultimately
we're
setting
up
in
the
lab
here
this
thermal
denaturation
assay,
where
we
put
compounds
in
and
then
we
thermally
denatured
to
look
for.
Broadly,
you
know
drug
binders,
but
that's
not
set
up
yet.
We've
got.
B
B
D
D
It's
really
complicated
when
I
have
the
pyridine,
the
in
the
fenugreek,
so
the
purification
is
getting
a
bit
harder,
but
after
seven
columns
sometimes
I
can
get
the
pure
compound.
So
I'm
gonna
work
in
this
compound
tomorrow
and
the
last
one
that
I
have
the
cf3
and
the
nitrile
group,
I'm
still
waiting
for
the
chemical
I
bought
from
apollo
is
on
demand.
D
D
So
after
the
reaction,
the
last
step
is
the
coupling
I
do
like
just
a
filtration
through
silica
light.
Then
I
go
for
normal
phase
using
chloroform
and
methanol.
Chloroforming
methanol
is
better
than
other
solvents,
so
I
try
the
other
ones.
Then,
for
this
specific
compound
I
use
the
column
the
amino
colon.
There
is
a
biotech,
a
minor
column.
We
have
here
it's
a
normal
phase.
So
after
that
I
went
for
the
reverse
phase,
the
reverse
phase.
I
used
methanol
and
ammonium
and
water.
D
So
I
got
the
salt
of
the
compound,
so
it
was
not
only
the
salt
was
a
mix
of
salt
of
the
compound
and
then
also
the
pure
compound.
So
the
nmr
was
a
bit
messy.
I
did
a
workup
yesterday
to
try
to
recover
it
from
the
water
phase,
but
so
far
it's
still
in
the
water
face.
Even
when
I
try
to
control
the
ph,
because
I
was
calculating
the
pka,
there
are
many
different
pks
because
I
can
get
protonation
in
three
different
nitrogens.
D
D
D
D
F
D
What
I
have
done,
it's
like
I
do
like
you,
methanol
and
water,
then
I
collect,
let's
say
10
troops,
so
these
10
tubes,
I
need
to
run
lcms
to
check
which
tool
I
have
the
pure
compound,
because
even
the
lcms,
sometimes
not
the
lsms
like
in
the
biostage.
It's
showing
me
just
one
one
signal,
but
then
in
the
lcms
some
tubes
are
mixed.
Sometimes
some
tubes
are
pure
and
some
tubes
are
only
the
final
compound.
So
it's
better
if
you
can
run
lcms
for
all
the
tubes
that
you
collect
from
the
body.
D
E
D
D
D
B
A
F
Okay,
so
these
are
the
analogs
on
which
I'm
working
on,
and
particularly
this
compound
should
be
ready.
I
have
to
to
check
the
purity
of
the
compound
and,
let's
see
if
the
nmr
is
okay,
but
this
looks
good
and
then
I'm
working
also
on
this
compound
and
I'm
in
the
middle
of
the
synthesis
of
the
reaction,
and
I
hope
to
to
have
also
this
compound.
E
Just
to
add
to
this
as
well,
we've
also
got
compounds
in
the
queue
for
the
the
nitrogen
scan
of
that
left-hand
piece
analogous
to
the
the
compound
in
the
dotted
box
with
the
purity
to
check
on
maria's
screen
maria.
Can
you
point
to
that?
Pyridine
ring
whoa
yeah
anyway,
so
those
are
coming
through
as
well.
B
Awesome,
I
think,
reading
availability
of
people
and
and
the
fact
that
we've
got
our
you
know
the
chocolate
holiday
coming
up.
I
think
it's
likely
we're
gonna
have
the
next
round
of
evaluation
as
towards
the
end
of
april.
I
would,
I
would
guess
so,
sometime
after
the
20th,
so
on
that
kind
of
timeline
yeah
I
mean
anything
that
you
can
ship
by
approximately
20th
or
something
is
likely
to
to
feed
in
nicely
to
the
next
round
of
evaluation
that
we
could
do
here.
E
I
did
actually
chase
up
wushi
for
the
missing
experimental
from
the
beginning
compounds,
so
I've
actually
met-
and
I
just
sent
you
the
document
so
that
you've
got
that.
So
we
have
that-
and
I
also
got
their
author
names
and
affiliations
for
the
publication.
It's
my
intention
to
read
that
this
week
I've
been
hoping
to
get
to
that
for
several
weeks,
but
anyway.
B
B
Great,
I
think
it's
important
we're
really
clear
about
that.
So
that's
that
sounds
fine.
All
right!
That's
wonderful!
Thank
you
for
chasing
that.
Okay!
If
there's
nothing
else,
then
that's
all
good.
We
will
likely
regroup
after
easter.
So
if
there's
nothing
else,
then
have
a
great
break
and
see
you
all
again
pretty
soon
bye,
then.
