►
From YouTube: DevoWorm (2021, Meeting 1): Overview of 2020, GSoC project preparation, and topical papers
Description
DevoWorm (2021, Meeting 1): Overview of 2020 presentation, review of caandidate GSoC projects, papers on condensates in Eukaryotic cells, Expansion Microscopy in C. elegans, and the first (Precambrian) embryos. Attendees: Susan Crawford-Young, Mainak Deb, Jesse Parent, Ujjwal Singh, Arnab Banerjee, and Bradly Alicea.
A
A
B
A
A
paper
that
we've
talked
about
over
the
past
several
meetings
in
the
previous
year
have
now
submitted
it
to
a
special
issue
of
biosystems,
but
we
also
have
the
preprint
up.
Here
is
a
open
access
version,
so
we
have
a
couple
of
what's
what
the
metrics
are
on
this,
so
I've
got
282
abstract
reads
since
about
friday.
I
think
it
was
accepted
on
friday.
A
A
A
A
A
Nlp,
okay,
well,
very
good
yeah,
it's
nice
to
see
everyone.
It's
always
fun
to
have
papers,
be
completed
and
released
into
the
wild
or
at
least
submitted
somewhere.
So
yeah,
that's
good!
So
again
we're
gonna!
I
think
we're
gonna
start
with
the
presentation
here,
and
this
again
is
just
kind
of
growing
into
2021.
This
is
going
to.
Let
me
share
my
screen.
A
B
A
And
so
there
are
a
lot
of
things
we
could
do
this
year.
In
fact,
last
year
I
was
going
over
the
presentation
for
last
year
and
I
found
that
a
lot
of
the
things
that
we
talked
about
doing
for
this
year.
We
kind
of
did
but
didn't
do
it
exactly
in
the
way
it
was
planned,
but
which
is
fine
and
we
did
a
lot
of
things
that
we
hadn't
planned
on
doing,
which
is
also
fine.
It's
actually
good
because
you,
you
know
when
opportunities
arise,
it
means
you
can
take
advantage
of
them.
A
A
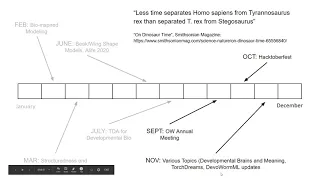
One
of
the
lessons
you
can
learn
from
this
paper
is
that
less
time
separates
homo
sapiens
from
tyrannosaurus
rex
than
separated
t-rex
from
stegosaurus,
meaning
that
the
tyrannosaurus
rex,
which
appeared
later
in
the
era
of
dinosaurs,
is
closer
to
us
today
than
t-rex
was
from
stegosaurus
or
a
dinosaur
that
appeared
early
in
the
dinosaur
era.
So
I
like
to
every
time
I
give
a
timeline.
A
I
like
to
give
a
quote
like
that,
to
put
the
flow
of
time
in
perspective,
and
so
thinking
about
like
how
things
happen
in
sequence,
you
know,
I
think
it's
enlightening,
but
for
our
purposes
we
have
2020,
which
may
have
seemed
a
lot
longer
than
it
was,
and
I
have
this
timeline
that
I've
prepared.
So
what
did
we
do
in
2020?
In
terms
of
our
meetings?
A
We
actually
did
a
lot.
There's
a.
I
have
a
spreadsheet
of
meeting
recordings
meeting
topics
and
dates
that
are
available
from
the
website.
If
you
go
to
the
weebly
site
for
diva
worm,
you
go
to
the
I
think,
the
last
the
final
tab.
It's
like
how
to
get
involved,
and
then
you
can
click
on
to
the
list
of
meetings.
A
For
2020,
and
so
all
those
meetings
are
listed
and
there
are
a
lot
of
really
interesting
topics,
but
two
of
the
topics,
two
of
the
things
we
did
like
in
february
and
march,
one
was
a
talk
by
tom
portages
who's.
A
a
member
of
the
group,
long-standing
member
of
the
group
b,
attends
the
meetings
occasionally
on
something
called
bio-inspired
modeling,
and
that
was
largely
on
a
lot
of
the
work
that
he
was
doing.
A
He's
been
doing.
He's
done
a
lot
of
work
with
something
called
morphozoic
he's
also
done.
Some
work
on
something
called
morphognostic,
which
is
a
sort
of
the
morphozoic
project
that
he's
working
on
involves
like
morphogenesis
morphognostic
involves
more
like
sort
of
cognitive
stuff,
and
so
currently,
I
think,
he's
modeling.
The
behavior
of
honeybees
he's
also
interested
in
nest,
building
in
fishes
and
other
types
of
things
like
that.
So
he's
got
a
lot
of
interesting
approaches
to
to
a
lot
of
interesting
problems,
and
we
had
him
give
a
talk
about
that
last
february.
A
We
also
had
a
talk
from
george
mikhailovsky
who's,
a
researcher
who
is
he
has
a
background
in
physics
and
biophysics,
and
he
did
a
talk
on
structuredness
and
entropy,
which
is
also
something
that
if
you
go
back
to
the
list,
you'll
see
it
on
the
list
in
march,
and
I
encourage
you
to
watch
that
video.
If
you
haven't
already
it's
based
on
a
paper
that
he
published
last
year
and
that's
something
that
we
you
might
want
to
revisit
hello
jesse.
How
are
you.
A
So
so
we've
had
the
two
meetings,
the
february
and
the
march
meeting,
and
those
are
just
topics
I
mean
there
are
a
lot
of
different
topics
in
the
meetings
if
you
forgot
or
if
you
didn't
make
a
lot
of
the
meetings,
I
encourage
you
to
go
over
that
list,
there's
a
lot
of
interesting
stuff
buried
in
there,
and
so
where
do
we
go
from
here
with
that
sort
of
stuff?
I
mean
that's
something
that
we
have
to
think
about.
B
A
A
A
You
know,
but
some
of
the
bigger
organizations
get
more
participants,
but
the
hacktoberfest
thing,
I
think,
is
maybe
something
we
revisit
this
year
with
a
little
bit
more
effort
into
it
or
you
know,
with
some
clear
goals,
because
we
just
kind
of
like
stumbled
into
it.
I
think
this
year,
but
I
think
you
know
I
think
it's
all
right.
I
don't
think
you
need
to
have
a
thousand
people
contributing
for
it
to
be
successful,
but
I
think
in
especially
for
the
diva
learn
platform
that
were
we
had
developed.
A
I
didn't
put
this
on
the
list,
but
around
the
same
time
we
released
this
diva
learn
platform
which
involved
the
different
google
summer
of
code
projects
and
I'll
talk
about
those
in
a
minute
but
yeah
we
released
it
around
here
and
then
we
got
into
hacktoberfest
almost
immediately.
So
I
think
next
year
maybe
we'll
have
some
clear
goals
in
terms
of
what
that
is,
and
then
in
november
we
covered
various
topics,
developmental
brains
and
meaning
torch
dreams,
diva
or
memo
updates.
A
A
Okay,
yeah
as
well
says
a
thousand
people
contributing
only
create
mess.
To
be
honest,
that's
probably
true
yeah
I
mean,
unless
you
have
the
infrastructure
for
dealing
with
that
many
contributors,
it's
hard
to
to
sort
out
so
yeah.
I
agree,
probably
in
this
case,
like
growing
oktoberfest,
would
probably
be
in
the
order
of
a
hundred,
but
yeah
I
mean
we.
We
can
plan
that
out
a
bit
more
and
I
think
you
know
the
important
thing
is
that
people
were
become
aware
of
it
and
that
they're,
you
know
excited
by
it.
Then.
A
Certainly
this
year
we
talked
about
or
we've
been
kind
of
working
on
this
divolent
paper.
So
I'm
not
going
to
cover
that
in
this
slide,
but
let
me
get
back
into
the
slides
here.
So
those
are
the
things
that
happened
during
this
year
and
there
are
many
more
things
that
happened.
I
didn't
put
them
on
here,
but
I
just
wanted
to
give
people
a
taste,
a
reminders
of
what
we
have
done.
A
A
A
E
A
E
A
A
Then,
in
september,
of
course,
I
did
the
diva
room,
2020
update
at
the
open
or
manual
meeting
and
again,
like
I
said,
I
think,
the
I
think
I
got
a
lot
of
people
sort
of
looking
again
at
or
at
diva
worm,
because
I
think,
like
people,
a
lot
of
people
in
open
realize
what's
going
on
in
evil
worm,
and
so
hopefully
you
know
that's
something
that
people
are
paying
attention
to
now,
and
certainly
it
was.
It
was
impressive.
A
A
This
was
by
krishna
katyal
and
myself.
We
worked
on
this
presentation,
so
basically
the
idea
was
to
talk
about
biological
neural
networks,
which
are
you
know,
networks
of
neurons
in
the
brain
and
then
artificial,
neural
networks,
which
are,
of
course,
you
know,
network
computational
networks
in
a
computer,
and
so
we
talked
about
the
differences
between
them.
You
know
talked
about
maybe
like
some
of
the
properties
of
neural
networks
that
we
can
exploit
that
are
more
biological
in
nature,
and
you
know
it
was
in
there.
A
A
So
I
mean,
if
you
wanna,
I
I
can
repost
the
blog
post
in
the
slack
channel
and
that
blog
post
is
everything
that
was
presented
at
neuromatch
three
with
the
video
link
and
the
slides
and
everything.
So
the
second
talk,
then,
was
the
bio
psychophysics
of
non-renal
cognition,
which
was
something
that
we're
doing
with
the
basil
area.
A
You
know
we're
focusing
on
these
diatoms,
this
diatom
genus
called
basal
area
and
we're
looking
at
how
they
move
in
in
the
water
column,
because
they're
diatoms,
they're
marine
organisms
and
so
we're
looking
at
actually
like
the
psychophysics
of
these
organisms.
How
do
they
move
and
maybe
how
does
how
do
you
stimulate
the
environment,
affect
their
movement?
So
that
was
a
talk
that
actually
went
pretty
well,
I
thought
that
it
was
well
received.
A
A
So
then
you
know
we'll
we'll.
You
know
we'll
wait
until
the
closer
to
the
deadline
to
sort
of
get
that
all
in
order
and
submit
it.
But
I
think
that'll
be
that'll,
be
good.
You
know,
that'll
help
us
follow
up
on
some
of
these
topics
that
we
did
public
lectures
on
in
2020,
publication
wise.
We
have
this
towards
the
digital
diatom,
so
this
was
published
in
diatom,
gliding
motility.
A
A
A
That's
another
follow-up
on
some
of
the
work
that
we've
been
doing
on
the
developmental
connectome
and
then
this
paper
data
theoretic
synthesis
of
the
early
developmental
process
in
december
2020.
This
was
accepted
to
neuroinformatics
on
in
their
special
issue
neural
commons.
So
we
this
was
a
paper
that
came
out
of
the
actually
it
it's
origin.
A
A
A
I
just
did
the
proofs
for
it
this
weekend,
so
it
shouldn't
be
too
long
before
it's
out
and
then
this
new
paper
periodicity
in
the
embryo,
which
again
this
is
something
usual
and
all
your
authors
on,
and
this
is
out
and
actually
borrow
my
archive
today
or
friday.
I
didn't
add
the
link
to
that
and
it's
also
submitted
to
biosystems.
A
Yes,
yeah,
it's
a
it's
a
pre-print
version,
so
I
mean
we'll,
probably
update
it.
You
know
with
a
new
version
if
we
have
comments
from
reviewers,
but
it's
it
yeah.
I
actually
can
put
the
chat
in
the
link
in
a
minute,
and
so
then
that's
good.
We
can
follow
up
on
those
publications.
Another
thing
I
didn't
add
because
it
isn't
really
out
yet
in
any
way
is
a
publication
on
the
on
the
devil
learn
material.
A
So
we
did
google
summer
of
code
2020
and
we
had
bojol
and
mayuk
and
they
were
our
students
for
last
year
and
they
worked
on
different
product.
My
oak
worked
on
this
project
evolearn,
which
was
a
pre-trained
model
for
microscopy
data
in
development,
and
it
came
out
very
well.
He
did
a
very
good
job.
He
set
up
this
pipey
repository
for
the
software
did
ai
project,
so
he
worked
on
bringing
together
a
lot
of
the
stuff
that
we
had
in
terms
of
ai.
A
In
terms
of
our
we
have
this
thing
called
devozu
that
has
a
lot
of
educational
materials
such
as
bringing
that
all
together,
and
so
they
both
did
very
well
in
their
projects
and
we
decided
to
build
a
github
organization
called
diva
learn.
So
this
is
the
software
diva
learn
and
then
there's
a
a
broader
github
organization,
called
diva,
learn
and
even
learn.
Involve
you
know
it.
A
A
So
we
released
this
thing
after
summer
of
code,
which
was
in
september
and
then
in
october
we
sponsored
a
hacktoberfest,
so
people
could
contribute
to
the
diva
learn
platform
through
oktoberfest
and
we
got
about
15
people
to
apply
and
and
to
to
participate
in
hacktoberfest,
which
is
you
know,
a
fair
number
it.
We
could
have
more
people,
of
course,
as
ojawa
mentioned,
if
you
scale
it
up
too
much,
you
know
we
really
do.
It
is
messy.
A
You
do
need
to
have
a
lot
of
people
that
dedicated
to
it,
so
you
know
we
might
want
to
grow
at
this
coming
year,
but
maybe
not
as
much
as
you
know,
some
other
organizations
that
have
more
more
attention
paid
to
it.
So
that's
something
to
think
about.
So
this
is
something
that
we
did
google
summer
of
code.
It
resulted
in
diva,
learn
and
now
you
know,
we've
had
public
events
based
on
diva
learn,
but
we'll
also
have
a
paper
coming
out
on
evil
learn.
A
Sometime
early
this
year
we
already
worked
out
a
paper
for
another
venue
that
didn't
happen,
so
that
paper
is
just
going
to
be
continuing
on,
and
I
I
hope
to
get
it
like
in
in
the
format
of
where
we
have
all
the
different
components
of
the
diva
learn,
platform
and
kind
of
give
a
good
good
description
and
then
release
it
as
a
preprint
as
a
good
first
draft,
and
then
we
can
maybe
publish
it
from
there.
So
I
think
that's,
I
think,
that's
moving
along
well.
A
This
is
diva
learn.
This
is
actually
the
launch
september
2020..
This
is
our
logo.
The
organization
is
on
github.
It's
github.com
divalern,
please
visit
and
check
it
out.
If
you
haven't
already.
This
is
our
weekly
meeting
youtube
archive.
This
is
another
thing
we
did
in
2020
and
this
is
you
know
I
mean
I
I
look
back
at
this
and
I
say
we
did
a
lot
of
stuff
in
our
meetings
and
we
have
a
lot
of
video
for
it.
A
So
now
the
question
is
is
like:
how
can
we
follow
up
on
some
of
the
stuff
because,
again
like
in
the
first
slides
I
mentioned
that
we
were
going
to
have
like
you
know,
we
had
all
these
things
that
we
did
different
points
in
the
air
and
it
kind
of
blurs
together
by
the
end
of
the
year.
So
you
know,
maybe
we
should
go
back
through
these
talks
and
look
at
like
some
of
the
things
that
were
mentioned.
A
A
A
A
A
Okay,
all
right
all
right
here
we
go,
let's
see
yeah,
let's
go
to
the
chat
working
on
it.
I
know
it
is
the
wait
a
bit
we'll
submit
soon.
That's
I
think,
on
the
one
of
the
papers
or
yeah
I
mean
yeah,
I
mean
we're
always
just
yeah
the
jaws.
Well,
the
jaws
paper
is
now
just
something
that
we're
doing
as
a
pre-print
like
the
just
just
people
thought
you
know
we
weren't
really.
A
You
know
it
wasn't
very
important
yet
in
terms
of
software,
so
they're
looking
for,
I
think
scientific
software
with
a
lot
a
pretty
big
user
base
so
but
we'll
be
working
on
that
paper
for
a
preprint,
so
yeah
well,
we'll
just
call
it
the
jaws
paper
as
a
as
a
shorthand
jesse
says,
I'm
curious
what
papers
are
in
development
now
or
can
be
contributing
to
yeah,
so
I
mean
I
think
we
mentioned
this
about.
Let
me
see
if
I
have
it
here.
A
B
A
Papers
and
again
this
isn't
updated.
I
need
to
update
it
more.
Some
of
this
stuff
is
like
kind
of
outdated,
but
these
are
you
know.
These
are
the
open
papers.
I
probably
will
put
in
something
on
the
on
the
diva
learn
paper,
although
that's
that's
just
really
kind
of
the
people
who
are
involved
in
setting
up
divo
learn,
but
people
can
contribute
if
they
want.
A
A
I
have
another
spreadsheet
in
another
place
where
we
have
submissions
that
are
going
to
be
made
and
I'll
put
those
up
on
this
readme.
Basically,
we
have
upcoming
papers
and
upcoming
venues,
and
I
put
it
in
a
document.
That's
I
don't
have
right
now
open.
I
can
actually
share
it
with
the
group,
maybe
in
in
the
open
or
in
the
diva
room
channel
and
the
open
worm
slack
as
well.
We
have
some
things
that
are
going
on
that
aren't
on
that
open
papers.
A
List
like
there
is
this
abstract
on
sort
of
developmental
biology
and
machine
learning
and
deep
learning,
but
that's
something
that
again,
I
I'll
probably
just
put
in
that
I'll
put
in
that
list
and
maybe
I'll
make
it
open
at
some
point
but
but
yeah.
I
think.
If
you're
interested
you
know,
maybe
we
should
talk
on
slack
about
it
like
what
specifically,
what
paper
specifically
are
you
interested
in
contributing,
because
I
mean
you
know
contributing
is-
is
a
kind
of
a
investment
of
time.
A
It's
like
deciding
what's
the
best
place
to
contribute
and
what
exactly
you
want
to
contribute
to
it.
Otherwise,
it's
kind
of
like
you're,
you
know
you
never
really
get
around
to
it
because
it
just
gets
overwhelming,
but
I
you
know
I
I
and
again
it's
like
things
that
are
sometimes
there
are
papers
that
have
been
sitting
there
for
a
while
and
they
just
need
some
focused
contribution
to
push
it
forward.
So
that's
another
thing
is
yeah,
so
I'll
go
back
over
this
list
again
and
try
to
update
it,
but
that's
where
it
is.
A
C
A
A
Yeah
yeah
the
periodicity
paper
is
like
this
is
the
first
draft,
so
we'll
be
doing
we'll,
be
getting
reviewer
comments
back.
So
there's
still
time
to
contribute
to
that.
You
know
it's
just
a
matter
of
like
you
know
it
can
be.
You
could
become
an
author
on
it
in
a
later
version.
So
that's
not
like
impossible.
It's
just
that,
like
you
know,
we
get
it
to
a
certain
point.
A
We
put
it
into
a
preprint
form
and
then
we'll
have
different
versions,
we'll
update
it
and
then
eventually,
if
it's
in
a
publication,
then
that's
considered
sort
of
the
end
of
the
road
for
that
and
then,
if
we
want
to
do
something
more
from
that,
we
can
write
a
new
paper
and
start
over.
So
it's
you
know
it's
that
sort
of
pr
iterative
process
where
we
build
up
to
this
publication
and
then
we,
maybe
you
know
if
we
want
to
do
we
find
something.
That's
interesting.
A
We
might
start
a
new
publication
from
that
or
you
know,
even
if
it's
something
that
doesn't
fit
into
an
existing
paper,
you
know
it's
something
we
could
follow
up
on
separately.
I
mean
that's
a
lot
of
things.
Start
you
get
like
you
know
they
butt
off.
You
know
you
think.
Well,
this
is
a
good
kernel
of
an
idea.
Let's
do
a
paper
on
this
or
let's
do
a
presentation.
A
You
know
they
they
kind
of
come
out
of
these
ideas,
so
I
mean
I've
written
paper
outlines
before
or
papers
where
they
just
kind
of
they
go
all
over
the
place.
They
don't
really
work,
but
we'll
take
a
couple
of
ideas
out
of
it
and
like
develop
them,
and
then
they
end
up
being
the
papers
that
are
actually
published
or
pre-printed
or
whatever.
A
So
I
mean
like
the
the
stuff.
I've
done
a
lot
of
stuff
with
we've
done
a
lot
of
stuff.
Earlier
in
this
in
diva
worm,
where
you
know
we
were
batting
around
ideas
and
and
developing
papers
and
then
you
know
there'd
be
some
ideas
that
would
come
out
of
it
like
the
networks,
the
embryo
network,
stuff.
You
know
that
kind
of
came
out
of
sort
of
applying
network
analysis
to
you,
know,
development
and
thinking
about
it
and
then
eventually
we
came
up
with
that
idea.
So
you
know
it's.
A
A
E
A
That's
what
I
understand
too
right,
yeah,
it's
10
weeks,
so
yeah.
Actually
that's
what
we're
going
to
do
next,
let
me
share
my
screen.
We've
got
so
we've
solicited
a
couple
of
ideas
here
from
people
over
the
holiday
and
again
so
thank
you
enough
for
attending,
and
yes
so
the
we
have
coming
up.
I
think
this
week,
incf
malen
from
incf.
A
A
A
So
this
is
like
so
there's
the
diva
learn
software
itself
and
then
there's
the
platform
and
I
think,
he's
interested
in
maybe
more
of
the
platform,
but
we
can
also.
There
are
other
components
to
this
for
them
to
work
on
as
well.
So
there
there's
the
divo,
learn
pre-trained
model,
there's
also
another
c
elegan
specific
platform.
That
was
the
product
of
an
earlier
gsoc
there's
some
other
software
that
we're
developing
in
there
in
terms
of
we
have
other
other
programs
that
we're
interested
in.
A
So
these
are
all
kind
of
fall
under
that
umbrella.
There
would
be
four
key
elements
in
this
project:
improving
the
current
models,
training
and
adding
more
useful
models.
So,
as
I
said,
we
have
this
group
of
models
already
that
we,
you
know,
maybe
want
to
improve
upon
or
add
new
models
or
modify
them
improving
usability,
which
is
always
a
good
thing
and
interactive
online
demos,
which
would
be
an
optional
thing
now,
with
10
weeks.
A
Of
course,
we'd
have
probably
have
to
picks
one
of
these
elements,
although
there'd
be
like
you
know,
if
we
picked
improving
the
current
models,
we
could
say
training
and
adding
more
useful
models
might
be
like
you.
You
would
work
with.
You
know
improving
some
of
the
models
and
then
proposing
next
steps
or
something
like
that:
improving
the
current
models.
We
have
the
basically
it's
like
benchmarking,
these
models,
so
using
a
number
of
methods
for
that.
A
So
that's
that's
one
thing
that
could
be
done:
adding
more
models,
so
diva
learn
should
also
contain
pre-trained
deep
learning
models
from
other
species
which
are
of
high
importance
in
developmental
biology.
We
can
split
up
the
different
species
as
so.
He
gives
an
example
here
in
pseudocode
for
developing
c
elegans
for
naval
and
zebrafish.
A
A
A
He
had
a
number
of
co-lab
notebooks
and
I
see
that
my
knock
actually
has
created
a
collab
notebook
as
well
on
some
of
the
stuff
he's
been
doing
that
as
well.
So
these
are
very
good.
I
I
like
collab
notebooks
because
they
give
people
nice
demos
in
a
very
you,
know,
usable
space.
You
know
people
can
just
go
to
the
collab
notebook
and
run
it,
and
so,
but
they're
still
a
bit
intimidating
for
people
from
non-cs
backgrounds,
so
we
should
focus
on
making
interactive
demos.
So
we
talked
about
this
a
bit.
A
So
yeah,
that's
that's
one
project.
Now
that's
proposed
that
now.
If
you
have
ideas,
you
can
issue
a
pull
request.
Maybe
you
know
ways
that
we
can
make
this
smaller,
because
this
has
to
be
fit
within
10
weeks.
So
this
is
a
lot
of
stuff
here,
and
this
could
be
three
or
four
projects,
but
we'll
probably
distill
this
down
to
like
maybe
one
and
then
you
know,
maybe
we'll
get
people
involved
in
other
things.
A
B
A
A
A
It'll
be
a
great
opportunity
to
learn
and
develop
a
cool
project.
We
need
documentation,
we
need
people
to
do.
You
know,
use
collab
or
jupiter
notebooks,
and
you
know
this
is
just
the
description
and
then
the
idea
would
be
to
create
sort
of
a
online
gui
that
would
execute
this
model,
and
so
that's
that's
the
vision
there
I'm
going
to
edit
it
some
more
and
we'll
probably
also
submit
this
one.
I
think
this
one
might
be
if
we
can
like
get
down
the
goals
of
what
we
want
them
to
do
in
10
weeks.
A
A
Okay,
so
usual
says
I
can
give
weekly
goals
for
the
project
yeah.
That
would
be
good
well,
if
we
could
like
have
some
sort
of
like
if
we
could
take
this
proposal
as
well
and
break
it
down
into
maybe
like
what
we
might
expect
over
the
for
you
know
every
two
or
three
weeks.
What
do
we
expect
from
them
and
then
then
we
can
like
get
it
because
I
mean
this
week.
I
just
need
to
know
the
like.
A
I
just
need
a
short
description
to
send
the
mail
one,
but
I
think
for
the
final
thing
that
we're
going
to
give
to
students,
I
think
it
would
be
good
to
have
like
you
know,
it's
always
good
to
give
them
like
kind
of
a
rough
outline,
because
they
have
to
build
those
schedules.
They
have
to
schedule
their
time
and
it's
always
that's
always
the
heart.
One
of
the
hardest
parts
of
the
proposal
is
making
a
timeline.
So
if
we
can
help
them
with
that,
that's
good.
G
A
A
Also,
it's
like
you
know:
it's
always
they've
been
they've,
been
interested
in
divorce
projects
and
they've
always
been
kind
of
tenuous
in
that
way,
because
you
know
we're
not
real,
I
mean
we're
open
worm
and
we're
you
know,
there's
we're
also
in
c
elegans
you're
also
segmenting
neuronal
cells,
but
we're
doing
developmental
stuff.
So
I
mean
I'll
just
I'll
just
put
a
statement
in
about
like
how
it's
sort
of
related
to
the
brain,
and
I
think
this
one
is
from
krishna.
This
is
devon,
specialized
convolutional,
neural
networks
for
biological
data.
A
A
A
Think
a
bit
maybe
a
bit
too
ambitious,
but
the
idea
is
really
different
than
what
most
people
were
doing
like
at
the
species
specific
level.
I
you
know
again,
we
haven't
really
done
anything
with
this,
so
it
might
not
make
any
sense
to
go
that
broadly,
you
know.
Maybe
it
makes
more
sense
just
to
look
at
you
know
across
species
instead
of
within
species,
because
the
models
that
we
have
now
are
really
kind
of
specialized
for
specific
species.
A
That's
the
way
we
kind
of
train
them,
so
their
performance
is
best,
maybe
with
one
species
or
another.
In
this
case
it
would
have
something
that
would
be
cross
species,
and
so
he
kind
of
lays
out
the
description
of
this.
This
is
again
something
we
need
to
work
on
a
bit
more,
but
this
is
another
idea,
so
we
have
those
three
ideas
and
I
think
that
those
are
probably
pretty
good.
A
I
don't
think
I
mean
we'll
probably
do
three
we'll
propose
three
projects,
and
then
you
know
we
will,
you
know,
maybe
get
students
for
all
of
them,
maybe
for
one
of
them,
maybe
even
for
none
of
them.
I
don't
know
what
the
situation
is.
This
year,
we
usually
get
a
lot
of
interest
in
the
machine
learning
projects.
A
F
F
A
A
F
F
A
A
Well,
it's
yeah.
It's
definitely
something!
Let
me
let
me
write.
Actually
I
could
write
that
up
myself
and
then
you
know
we
could
actually
submit
that
as
well.
Yeah.
So
that's
that's
another
one
and
then
that's
something
we
can
do
like
outside
of
g
sock
as
well,
but
yeah,
but
thanks
yeah,
that's
good!
That
sounds
good.
A
A
D
A
Well,
that
sounds
good
yeah.
That
sounds
good.
Look
forward
to
the
pull
request.
We
also
have
three
papers
here:
I'm
gonna
go
through
them
really
quickly.
If
you
have
to
leave
that's
okay,
but
I
mean
I
just
wanted
to
go
through
these
things
that
I
found
here
or
actually
well.
The
first
one
we'll
go
through
here
is
the.
A
F
A
F
F
A
Yeah,
so
this
is
a
picture
of
the
nucleolus.
The
larger
largest
structure
inside
the
nucleus
is
a
condensate
with
internal
structure
and
the
steam
nuclei
from
frog
cells.
Condensates
of
different
proteins
nest
with
one
another.
So
a
condensate
is
a
sort
of
a
physical
property
of
something
where
there's
a.
A
G
F
A
D
A
So
I
mean
you
know
these:
these
quantum
articles
they
kind
of
there's
they
have
a
narrative,
so
it's,
but
it's
pretty
easy
to
follow
along,
and
so
they
they
were
talking
about.
Let's
see,
they
were
talking
about
the
phase
separation
in
cells
trickled
up
around
2012..
I
wonder
if
this
hymen
is
the
same
person
who
did
the
there's
a
method
that
I
used
to
use
for
getting
different
fractions
of
rna
out
of
a
cell?
I
wonder
if
I
don't
know
if
this
is
the
same
person,
but
it
sounds
like
it.
A
That's
what
maybe
they're
trying
to
do
here,
they're
trying
to
like
figure
out
different
parts
of
the
face
separating
different
parts
of
the
cell
out
and
so
biomolecular
condensates
in
cells
in
complex
cells,
phase
separation
drives
proteins
and
other
molecules
to
aggregate
into
fluid
bodies
or
condensates
with
diverse
functions.
A
few
of
these
condensates
are
shown
here.
So
this
is
different
things
like
nuclear
speckles.
A
A
You
know
condensates
and
gene
expression,
so
condensate
seem
to
be
involved
in
many
aspects
of
cellular
biology,
but
gene
expression
and
the
production
of
proteins
is
one
area.
They
talk
about
the
ribosome,
so
this
is
yeah.
There's
some
interesting
stuff
there.
The
ribosome,
of
course,
is
where
rnas
get
translated
into
amino
acids,
which
then
build
proteins.
A
F
F
A
B
A
So
all
these
things
too,
you
know
they
have
like
if
you're
interested
in
machine
learning,
I
mean
there's
a
lot
of
that
that
if
we
can
get
images
and
people
are
producing
image,
high
resolution
images
they've
also
produced.
You
know
like
of
diff
these
different
parts
of
the
cell,
there's
a
lot
of
em
or
electron
microscopy
data
out
there.
A
A
So
it's
like
one
way
to
get
at
very
small
structures
I
mean
you
can
one
way
is
to
get
really
high
resolution
microscopy.
Another
way
is
to
expand
things
out,
and
so
at
least
as
I
understand
it,
I'm
not
I'm
not
an
expert
at
this.
So
but
they've
done
this.
In
c
elegans
specifically,
so
this
is
the
paper
describing
this
technique
applied
to
c
elegans,
so
they've.
A
So
thus
we
set
out
to
develop
this
protocol
customized
for
c
elegans,
so
they
had
to
work
out
a
custom
protocol
for
this
because
of
c
elegans
biology
makes
it
hard
to
do
this,
so
they
did
this
see
if
they
have
any
images
this
well.
This
is
a
diagram
of
what
they
did
for
the
protocol
and
then
I
wondered
if
they
actually
had
any
microscopy
images
in
here.
A
A
B
A
Is
the
correct
or
sort
of
a
popular
term
in
places
where
that
is
bound
to
the
the
where
that
fluorescent
molecule
ends
up
by
and
gone
bound
to
the
target?
So
you
can
see
like
in
this
area
with
red,
you
have
a
lot
of
this
fluorescent
molecule
bound
to
its
target
and
the
places
where
it's
not
red,
it's
not
and
so,
and
they
show
like
kind
of
examples
of
this,
where
they're
able
to
do
this
with
different
methods.
A
So
this
is
you
know
it's
one
example.
I
don't
know
if
you
can
see
it
very
well,
so
this
is
another
one
where
they.
This
is
a
better
actual
example.
You
have
liberal
stages,
two
through
four
or
two
four
and
then
day
one
adult.
So
these
are
different.
This
is,
as
the
cell
is
developing
post
embryonically.
A
You
have
pre-excel,
which
is
before
they
implement
the
method
post
excel,
where
it's
expanded
by
3.3
times
its
original
size
and
then
they're
they're
doing
this
registration
technique
where
they're
overlaying,
the
pre
and
post
excel
images,
and
so
here
where
they
have
like
basically
a
worm
where
they
haven't
applied
this
technique,
then
they
have
applied.
The
technique
and
then
they're
registering
the
image
so
that
they
can,
you
know,
figure
out
where
things
are
in
the
image
and
right.
So
that's
that's
an
example
there.
A
A
A
In
the
distilled
water,
but
I
think
yeah,
I
think
the
idea
is
they
can
do
this
in
a
controlled
manner,
because
you
can
imagine
like
if
you
expand
a
bunch
of
cells
like
if
you
freeze
an
organism
and
you
unfreeze
it
a
lot
of
times.
There's
a
lot
of
tissue
damage
like
in
c
elegans
specimens.
If
you
freeze
down
you
can
freeze
down
c
elegans
like
in
a
you
know,
you
can
freeze
them
in
a
solution
and
then
bring
them
back.
A
Hydrate
them
from
you
know
they
thought
thaw
them
out
and
hydrate
them,
and
you
know
there'll
be
some
eggs
that
survive
and
you
know,
but
there's
a
lot
of
tissue
damage
to
the
worms
that
were
already
in
that
sample.
So
you
know
you
do
you
have
to
do
this
in
a
controlled
manner?
I
guess
is
my
point,
and
so
this
is
showing
kind
of
how
this
method
reveals
anatomical
structures.
A
So
you
know
it's.
Some
of
these
anatomical
structures
are
very
clear,
much
clearer
than
you
would
get
with
something
like
like
a
standard
microscopy
technique,
but
there
is
this
electro
electron
microscopy
technique,
that's
better,
but
it's
harder
to
use,
and
so
this
is
just
using
like
a
standard
microscopy
setup
but
you're
actually
getting
a
higher
resolution
of
things.
A
I
don't
know
how
it
compares
to
electron
microscopy,
but
these
are
pretty
good
images,
so
I
don't
know
so
yeah
this
paper
again.
If
you
want
to
read
this
paper,
it's
a
protocol,
so
it's
probably
gonna
be
hard
to
read
for
many
of
you,
but
just
to
go
through
the
images
just
to
get
an
appreciation
of
the
technique.
A
So
jesse
says
I
know
logan
thrasher,
thresher
collins
is
into
this
stuff
and
ed's
work.
If
there's
a
particular
need,
I
could
ask
him
to
come
to
the
group
and
talk
some
time.
Yeah
that'd
be
actually
good.
If
you
could
set
that
up.
That
would
be
a
good
talk
to
have,
I
think,
would
be
a
lot
of
interest
in
it
so
yeah.
A
A
Many
embryos
are
an
early
cleavage
stage
and
most
of
them
yield
of
limited
phylogenetic
signal,
which
means
that
they
find
embryos,
but
they
don't
exactly
know
where
they
fit
into
the
tree
of
life.
So
this
is
just
a
report.
This
is
a
standard
like
paleontological
report
on
what
they
found,
and
so
here
we
report
the
discovery
of
two
embryos
that
are
three-dimensionally,
preserved
and
complex,
imaging
techniques
using
propagation
phase
contrast
based
synchrotron
radiation.
A
Unexpectedly,
our
observations
show
notable
noticeable
difference
in
organization
patterns
between
the
embryos
suggesting
that
they
represent
two
distinct
taxa.
That
means
that
they
have
they're
of
two
different
species,
and
so
these
are
they're.
Comparing
embryos
and
they're,
saying
that
there
is
more
than
one
species
in
this
in
this
assemblage.
A
These
embryos
provide
further
evidence
of
for
the
presence
of
bilitarian
animals
in
the
in
this
biota,
which
is
this
group
of
of
things
that
they've
found.
Furthermore,
these
bioitarians
have
already
diverged
into
the
distantly
related
groups,
at
least
40
million
years
before
the
cambrian
radiation,
which
is
a
time
in
evolution
where
you
start
to
get
a
lot
of
different
species
like
the
birth
of
sort
of
vertebrates.
A
Vertebrate
life
fishes
all
of
that
come
in
the
cambrian
or
in
in
this
radiation,
indicating
that
the
last
common
ancestor
of
the
biletarians
lived
much
earlier
than
is
usually
thought.
So
basically,
this
paper
tells
us
that
embryos
existed
before
the
cambrian
explosion,
which
is
a
time
when
we
thought
that,
like
a
lot
of
vertebrate
life
originated-
and
there
was
before
this
this
time
of
vast
radiation-
it's
really
not
clear
what
was
there.
A
We
know
that
there
were
some
microbes
and
some
other
organisms
that
that
existed
back
then,
but
we
really
didn't
know
anything
about
when
embryos
emerged,
and
so
what
they're
suggesting
is
that
embryos
emerged
much
earlier
than
a
lot
of
the
a
lot
of
the
early.
You
know
a
lot
of
the
life
that
emerged
in
the
cambrian
explosion,
and
so
this
is
pre-cambrian
post-gastrular
embryo.
This
is
the
example
of
the
embryo
here.
A
A
A
I
don't
know
if
that
means
I'm
not
reading
enough
for
what,
but,
but
you
know
this
is
a
very
interesting
paper.
It
kind
of
goes
on
about
this
embryo,
and
this
is
really
the
first
embryo
that
we
found
again
it's
you
know.
This
is
a
time
when
you
had
sort
of
the
emergence
of
what
they
call
medicines
which
are
animals.
A
You
don't
really
have
any
distinction
between.
You
know
you
have
bilaterians,
which
means
they
have
the
two
sides
of
the
you
know
the
left
side
and
the
right
side
of
the
organism
which
are
sort
of
mirror
images.
So
there's
that
symmetry.
But
there's
really
you
know
you,
you
have
like
not
much
else.
You
don't
really
have
vertebrates.
You
don't
have
invertebrates,
as
as
we
know
them
today,
they're
just
these
bilitarians
that
are
exist,
and
these
are
like
the
early
versions
of
these
biotarians
so
yeah.
This
is.
This
is
an
interesting
paper.
