►
Description
Open project meeting for Open Source Antibiotics Series 1, the Mur Ligases.
Full Project: https://github.com/opensourceantibiotics/murligase
Relevant GitHub Issue: https://github.com/opensourceantibiotics/murligase/issues/90
On the call: Professor Matthew Todd, Dr Edwin Tse, Yuhang Wang, Dr Daniel Gedder, Andy Kim (UCL), Dr Adrian Lloyd and Dr Laura Diaz Saez (University of Warwick), Dr Joe Eyermann, Dr Lori Ferrins (Northeastern University).
A
And
I
will
just
share
the
right
thing,
which
is
here:
whoops,
okay,
so
hopefully
people
saw
the
if
you
I
shared
and
it
outlines.
You
know
lots
and
lots
of
things
which
are
going
on
to
my
mind,
of
all
the
things
we're
doing
at
the
moment
which
are
listed
here.
The
the
things
I
want
to
talk
about
most
would
be
latest
results
from
from
you
guys,
Laura
and
Adrian,
and
also
just
comments
on
the
latest
structures
that
came
through
and
and
just
to
to.
A
You
know,
prefigure
that
perhaps
given
them
all
the
new
data
we've
got
recently,
we
might
want
to
have
another
dedicated
session
on
crystallography,
so
soak
in
number
two.
But
we
could
talk
about
the
new
structures
and
what
we
learned
from
those
and
what
we
need
next,
if
anything
from
the
SSC
CID
guys.
So
if,
if
you're
happy
to
the
Warwick
team
to
take
over
and
and
talk
about
what
you've
been
doing
recently,
that
will
be
I'm
sure
of
interest
to
everybody.
B
B
So
I've
been
doing
some
spr
the
past
two
weeks,
I
think
no.
Last
week
yeah
we
can
have
so
I've
got
some
results
for
SBR,
with
economyod
on
the
April
confirmation,
with
the
competition
compounds
for
this
first
round.
So
I've
got
some
KD
values,
but
take
them
with
a
pinch
of
salt
right
like
I
would
say.
This
is
a
hundred
meter,
more
100
I
would
say
these
are
very
similar
because
the
fitting
Is
Not
Great,
because
there's
more
data
points
to
add.
B
A
B
A
A
B
So
there
is
no
specific
language,
they
look
like
a
straight
line.
Basically
now
in
these
binding
right,
you
kind
of
determined
which
one
is
specific
and
which
one
is
not
a
specific.
It's
just
how
the
curves
are
looking.
It
might
be
a
mixture
of
both,
but
we
are
never
sure
about
that.
Does
that
would
be
with
any
binding
experiment?
Basically
right,
you
never
know
if
you're
buying
the
one
two
size
or
three
in
this
kind
of
things,
especially
with
low
binders,
that
you
cannot
discriminate
within
different
binding
sites.
B
A
B
B
B
D
B
B
Well,
not
at
the
level
not
so
I
cannot
find
the
final
hits
from
them
that
we
are
following
up.
The
best
hits
on
the
essay
I
can't
find
them
on
the
SBR
as
heads,
but
we're
gonna.
Look
after
I
do
the
experiments
on
the
door
response
and
I
make
sure
which
ones
are
not
specific
and
which
ones
are
specific.
C
C
C
There's
an
atom
wise,
Library
update,
so
This
concerns
the
compounds
that
did
inhibit
the
amplex
grad
base
coupling
system,
the
ones
that
we
in
the
initial
phase
have
to
discard.
Because
of
that,
these
15
compounds
were
being
screened
with
an
assay
that
follows
ADP,
however,
kinase
lactate
is
recognized
and
in
actual
fact
14
of
these
at
one
millimole
is
still
significantly
inhibited
the
coupling
system.
C
So
otherwise,
so
in
the
background,
a
number
of
things
have
been
happening,
for
example,
the
synthesis
of
substrates,
the
establishment
of
America
assay,
which
we've
done
and
we've
now
I've
got
to
process
the
data.
But
we
have
those
response
occurs
for
cediments
regional
summer
F
versus
our
control,
inhibitor
adpcp.
C
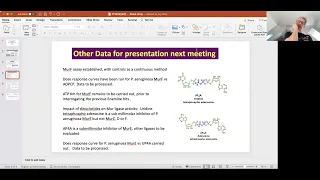
We
need
to
characterize
the
enzyme
kinetically
before
it
goes
into
the
enamine
screen,
and
finally,
I
mentioned
this
before
that
we
have
the
idea
that
molecules
like
eurodine
tetraphosphodenazine,
which
have
a
uracil
and
an
adenine
nicely
spaced
between
them
might
actually
impact
and
the
ligase
at
both
the
ATP
and
UDP
substrate
burning
site.
It
turns
out
that
up4a
is
of
all
the
ligazers
we've
tested
a
specific
inhibitor
from
mer
e.
A
A
Compounds
are
odd
right,
so
on
I
remember
the
from
the
atom-wise
Library
yeah
I
mean
the.
As
you
said,
there
were
15
compounds
that
appear
to
mess
up
the
the
coupling
enzymes
and
and
it's
the
same
issue
you're
talking
about
with
the
competition
compounds
right
that
they're
doing
the
same
thing.
Yeah.
C
C
A
C
To
be
honest,
I
don't
know
off
the
top
of
my
head,
I
I
don't
have,
we
would
have
to
look
I
mean
we
we,
we
could
actually
start
to
investigate
further
and
looked
at
the
impact
on
individual
other
enzymes.
I
mean
do
you
have
the
the
other
possibility
is
is
that
we
try
and
steer
away
we
steer
away
from
doing
coupled
assays.
In
the
same
the
way
we've
been
doing
for
these
compounds
and
maybe
look
at
some
of
the
chemical
methods
for
phosphate
release.
C
The
reason
we
haven't
done,
that
is
they.
Those
methods
require
use
of
a
fairly
High
concentration
of
acid
to
get
color
development
and
I
have
a
problem
with
that,
because
the
year
the
UDP
murloc
peptides
break
down
significantly
and
also
will
liberate
phosphates
under
those
circumstances.
So
that's
that's
a
can
of
worms
that
I
wanted
to
try
and
avoid
opening.
If
I
could
avoid
doing
so.
A
A
D
A
We
will
take
a
look
at
the
structures
for
interest
and
we'll
we'll
give
an
interim
report
that
we'll
make
it
clear
that
it's
work
in
progress.
Yeah
I
mean
it's
very
nice
that
there
is
apparently
something
there
from
spr
which
really
helps,
and
you
know
we
don't
know
where
it's
binding
or
what
it's
doing.
That's
encouraging.
A
A
A
B
B
D
A
A
A
A
D
A
Great,
so
this
is
a
work
in
progress
with
the
atomized
compound,
so
we're
going
to
cover
other
things
about
going
back
to
atomize
just
yet
until
we've
completed
the
data
all
right
just
on
on
the
shipping
of
compounds,
I
think
you
hang
has
finished
in
their
ship.
This
key
Amy
derivative
of
the
original
AZ
hit.
B
A
A
A
A
C
A
D
A
B
A
A
I
mean
this
will
be
stable,
no
problem,
it's
just
the
Amy,
that's
the
issue.
So
so
you
hang
is
now
now
the
chemistry
is
starting
to
behave
itself
and
the
purification
is
easier
he's
playing
on
other
diamines
here.
You
know
in
case
this.
This
Linker
length
is
wrong.
There
are
other
diamines
you
can
get
with
bot
groups
on
one
end
and
we
can
make
other
compounds
now
mm-hmm
all
right.
Great
thanks-
and
you
know
it's
great-
that
you've
got
a
sample
of
this
guy.
A
Okay,
so
just
to
switch
to
that,
because
that's
the
other
thing
that
we
got
updates
from
I
think
we
saw
these
yeah,
so
we
saw
the
these
have
been
done
last
time.
This
is
a
structures
from
yeah,
ssdcid
and
I.
Think
last
time
reported
that
these
were
were
done,
but
they
hadn't
been
deposited
so
now
they've
been
deposited
and
they
can
be
accessed
using
these
URLs
here,
which
is
great
and
the
ligands
that
are
involved
in
in
The
Binding
to
Mercy.
A
A
A
D
D
Yeah,
these
two,
your
currents,
are
showing
I'm
pretty
sure,
because
I
double
checked
with
pdb
yeah
and
with
the
paper
so
I
found
the
structures
exactly
matching
these
two.
The
problem
is:
are
we
sticking
to
the
AZ
company
numbering
or
are
we
expect
to
do
the
AZ
commercial
compound
number?
Because
we.
A
D
That
these
pay,
these
numbers
are
easy
to
track
because
I,
when
I
search
for
the
commercial,
for
example,
the
legend
AAC
one
three,
six,
four
or
whatever
it's
I
cannot
find.
I
cannot
find
any
website
resources
that
can
like
just
link
to
this
structure.
So
I
think.
Maybe
we
stick
to
the
paper
numbering
it's
a
bit
easier
for
us
to
identify.
A
A
Yeah
so
I
think
Joe
sent
us
a
file.
Wait
just
let
me
check
here
one
sec,
always
every
single
project.
You
know
you
have
issues
with
correlating
across.
You
know,
numbers
between
different
coding
systems,
so
Joe
yeah
kindly
sent
this
earlier,
and
so
this
is
the
content
that
we're
talking
about
so
become
a
26
in
the
paper
with
that
structure
which,
based
on
the
picture
that
Yan
sent,
looks,
looks
right.
A
So
they're
all
yeah
they're
all
got
the
same
they're
saying
the
same
core
and
different
side
chains,
which
is
great.
So
that
is
a
very
nice
set
of
compounds
with
multiple
compounds
bound
to
the
same
enzyme,
allowing
us
to
to
really
see
what
impact
the
side
chain
makes
if
anything,
but
they're,
all
bound
the
same
way.
I
think.
A
Okay,
so
if
that
is
the
yeah,
so
if
that's
correct,
then
you
hang
then
we're
all
good
based
on
the
structure
that
was
that
you've
put
up
here
and
so
there
it
is,
which
is
great.
Thank
you.
Ian
in
your
absence.
That's
really
beautiful
structures,
I
think
he's
going
to
finish
this
off
and
then
you
know
submit
it
as
per
usual.
E
D
E
A
A
A
D
E
But
I'll
try
to
get
the
Styles
off
first
and
keep
working
the
other
synthetic
roads,
because
if
you
go
a
little
bit
down
yeah
this
oxidizole
I
I
needed
to
change
the
first
step,
because
I
was
planning
to
do
a
reaction
with
asset
key
and
hydride
and
then
bramination
with
NBS,
but
didn't
work.
So
I
repeat
the
reaction
with
the
tube
bromo
and
hydride
it's
working.
E
However,
purification
is
a
bit
confused
now,
but
I
could
see.
I
have
this
compound
lcms
and
what
I'm
thinking
to
do
is
turned
off
synthesize
again:
the
Boost
enemy,
I'm,
gonna,
do
or
Gabriel
cities
or
whatever,
to
convert
the
bromine
to
I'm,
trying
to
use
the
outside
to
convert
bromine
to
the
online
and
then
I
can
use
the
quadruple
mild,
because
the
first
step
that
I
was
working
to
synthesize
the
dynamite.
E
They
yield
is
so
really
bad.
It's
like
five
percent
and
I'm
trying
to
do
this
step
first
and
then
I
can
go
to
the
dialysis
or
the
oxidizole
as
well.
Okay,
the
last
the
last
step
also
a
couple
of
problems.
The
first
did
the
work.
Then
I
tried
Suzuki
coupling
a
major
product
that
I
get
was
not
the
product
from
the
Suzuki
coupling.
What
I'm
doing
is
now
kind
of
protected
the
bromide,
the
reaction,
work
and
then
I'm
gonna.
E
A
A
D
A
Just
to
note
that,
as
we
get
more
of
the
more
certainty
on
the
inhibition
by
the
the
the
app
the
enemy
compounds
from
the
Warwick
Union
collection
and
the
atomized
compounds,
you
may
have
seen
that
Ed's
already
been
doing
some
assessment
of
near
neighbors
of
these.
So
you
know,
I
mean
what
we
really
want
to
do.
If,
if
we're
happy
with
some
of
these
hits
is
to
then
start
looking
at,
you
know
what
it
makes
most
sense
to
buy.
A
So
he's
already
started
looking
at
this
for
some
of
these
compounds
that
were
reported
as
being
multi-targeting
Inhibitors
and
just
extracting
the
obvious
compounds
that
we
can
buy.
That
would
flesh
out
some
SAR
and
obviously
it
makes
sense
to
do
that
if
we're
really
confident
about
the
fact
that
they
are
binding
and
doing
what
we
want.
A
It's
a
judgment
call
about
when
we
start
with
that
so
on.
The
left
here
are
the
ones
in
the
Warwick
collection
and
on
the
right
of
the
atomized
compounds.
It's
a
judgment
call
right
because
each
of
these
It's
relatively
quick
and
easy
to
get
enough
material
at
about
you,
know
100
euros
per
sample
or
something
and
in
the
grant
that
we've
got
from
antibiotics
research
UK
we
have
money
to
spend
on
this
stuff,
I
mean
particularly
on
the
otherwise
compounds
where
we
can
use
it
for
anything.
A
A
C
Yeah
I
mean
I
think
if
it
were
just
me,
I
think
I
probably
will
go
ahead.
To
be
honest,
I
think,
if
simply
because
I
know
that
they
actually
do
work
as
Inhibitors
and
and
they
multi-target
the
question
about
how
they're
doing
it
is
something
that
we
we
still
have
to
flesh
out.
But
nevertheless
they
do,
and
this
is
what
we've
in
principle
been
after.
A
That
will
be
good
yep
I
I
mean
I
agree,
because
these
came
from
larger
collections,
where
there
were
many
compounds
that
didn't
do
anything,
and
so
you
know
there's
plenty
of
negative
controls
right
of
compounds
that
were
part
of
the
same
screenings
that
didn't
didn't
do
anything
interesting.
So
it's
a
good
sign
that
we
should.
We
should
investigate,
compounds
more
and,
of
course,
if
we
do
start
playing
around
with
these
and
we
get
we
get.
C
A
A
Right,
good,
all
right
in
that
case,
thanks
for
coming
along
everybody
and
see
you
next
month
and
I'll,
send
out
some
possible
dates
for
for
a
focused
chrysologically
discussion.
Obviously
I'd
like
to
get
the
ssgc
ID
people
along
and
lesborough
along
and
make
sure
everyone's
involved
in
that
so
I'll
try
and
find
a
decent
time
for
that.
