►
Description
Open project meeting for Open Source Antibiotics Series 1, the Mur Ligases.
Full Project: https://github.com/opensourceantibiotics/murligase
Relevant GitHub Issue: https://github.com/opensourceantibiotics/murligase/issues/86
On the call: Professor Matthew Todd, Dr Edwin Tse, Yuhang Wang, Kato Leonard (UCL), Professor Chris Dowson, Laura Diaz Saez, Adrian Lloyd (University of Warwick), Lizbe Koekemoer (Diamond/Oxford), Dr Lori Ferrins, Dr Joe Eyermann (Northeastern University), Dr Chris Swain (Cambridge MedChem Consulting), Dr Jan Abendroth (UCB Biosciences), Dr Bart Staker (SSGCID).
A
Okay,
so
it's
the
12th
of
july-
and
this
is
an
open
source,
antibiotics
update
meeting
for
the
merly
gay
series,
and
I
do
I-
I
posted
a
very
sort
of
sketchy
issue.
Yesterday
the
recording
is
up
of
last
meeting
and
I
posted
a
very
sketchy
update
for
this
meeting.
Apologies
for
that,
but
I
I
just
took
the
most
important
things
and
and
put
them
up
there,
and
I
I
mean
we
can
take
these
in
any
order
and
and
and
deal
with
the
most
important
things.
A
First,
I
guess
from
from
my
perspective
the
two
things
that
we
should
start
with
would
be.
You
know,
as
as
a
result
of
the
the
exciting
data
that
were
presented
last
time
by
adrian,
whether
we
have
any
updates
on
on
that
work
from
adrian
and
laura
and
then
also
whether
we
have
any
updates
on
the
structural
biology.
A
I
think
that
yan
is
not
here,
but
whether
we
have
any
new
structures
that
we
should
be.
We
should
be
talking
about
because
I
think
that
there
were
lots
of
oh
yes
just
joined
there
were
there
were
lots
of
things
I
think
on
on
the
stove
as
it
were,
the
people
were
thinking
about
what
people
are
trying
it'd
be
great
to
catch
up,
so
laura
and
aj
are
both
here
and
ready.
Did
you
want
to
lead
off
yeah.
B
D
C
B
So
I've
been
doing
a
few
things
for
crystallization,
so
I
have
been
following
up.
They
have
been
following
up
the
main
hits
from
the
last
structures
for
equal
amino
d,
so
we
can
use
them
for
x-cam
and
soaking.
In
the
case
of
mudadp,
I
got
surprisingly
less
proportion
of
crystals
than
I
was
expecting.
I
was
expecting
it
to
be
more
successful.
B
Then,
in
the
case
of
emi
the
monopeptide,
I've
got
some
crystals
more
crystals
than
with
adp,
but
the
compounds
were
precipitating
in
the
crystallization
condition,
so
I
think
they
were
impairing
the
soaking.
Still
I've
got
some
crystals
soaked
and
they
are,
they
have
been
sent
to
diamond
and
also
I
want
to
try
another
condition
for
you
and
my
given
that
that
condition
was
precipitating
the
components.
B
Next,
I've
done,
cocky
sterilization
with
the
four
hits
the
top
hits.
We
are
working
on
and
those
so
they
I
don't
have
any
crystal
growth
yet,
but
there
are
still
more
things
that
I
can
try.
So
that's
gonna
start
in
round
two
of
the
crystallization
and
it
will
involve
three
proteins:
new
cd
and
e
with
the
compounds.
So
let's
see
how
that
goes
on
the
crystallization
e
coli.
D
B
B
C
B
B
Again,
all
that
is
going
to
diamond
solubility.
There
were
solubility
issues
with
the
compounds
again,
but
less
issues
than
in
the
case
of
the
crystals
and
yeah.
I
also
prepared
some
more
e-coli
crystals
for
the
close
conformation
in
with
different
concentrations
of
the
tree
peptide,
but
I
haven't
got
any
crystals
yet
and
I
also
started
cooker
salutation
with
pseudomonas
musi
and
monopeptide
and
again
I
got
no
crystals
yet
for
that.
B
E
E
B
B
So
I
started
with
millimolar
okay
and
then
I
did
the
illusions
as
well,
so
I
was
basically
based
on
the
percentage
of
them.
So
as
I
know
this
compound
these
stuff,
these
crystals
don't
go
more
than
ten
percent
of
the
color
of
the
dmso.
So
and
I
started
with
that
and
then
they
hit
it
down.
But
everything
was
the
same.
E
Okay,
I
just
thought:
maybe
you
know
if
you're
doing
high
millimolar
concentrations,
if
you
know
if
you
could
get
by
with
something
like
100
micromolar,
but
maybe
you
have
already
done
that
experiment.
You
said
you've
done
dilution.
So
if
you
get
a
10-fold
dilution,
I
assume
you
started
at
5,
millimolar,
1,
millimolar,
then
10-fold
dilution.
Then
you
would
be
down
in
that
yeah.
B
Actually
I
started
with
two
millimolar
because
one
of
them,
the
solubility,
was
already
not
that
great
in
the
first
dilution
that
I
did
and
with
100
myself.
So
I
just
have
to
see,
if
I
remember
correctly
but
again,
I
you
know,
I
want
to
check
again,
because
I
did
a
lot
of
things
that
day,
not
only
this
project.
Okay,.
B
D
I
I
just
wonder
laura
what
what
do
you
think
clearly,
this
structure,
it
sounds
like
there's
a
structural
change
going
on,
so
it
might
be
either
it
can
only
be
done
by
co-crystallization,
or
is
it
possible
to
do
some
ligand
exchange
interactions
at
the
binding
site
if
we
had
some
really
weak
analog
present
in
the
in
the
site
where
the
inhibitor
is?
Is
it
possible
to
do
some
kind
of?
Is
that
even
a
thing
where
you
can
kind
of
do
an
exchange,
so
you
can
kind
of
maintain
the
structure?
Somehow
that's
that
not
happened.
B
B
E
D
F
B
E
That's
great,
actually
laura
so
you're
just
making
the
comment
about
magnesium.
So
that's
that's
kind
of
a
little
tricky.
I
think
so.
If
you
look
at
the
structures
of
uma
adp
bound
with
magnesium
at
least
based
on
how
we're
thinking
some
of
these
compounds
might
be
binding,
magnesium
actually
may
be
blocking.
So
if
you
have
the
so,
I
was
thinking
out
if
you
did
any
experiments
with
uma
with
no
magnesium.
I
don't
know
if
that's
even
possible.
B
E
F
F
F
C
B
C
A
C
C
C
B
D
F
So
this
will
be
fairly
quick.
So
what
this
is?
This
essentially
is
finishing
off
a
section
of
work
which
I
was
basically
gave
the
presentation
last
time
about.
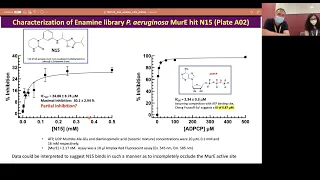
So
this
concerns
pseudomonas
original
summer
e
and
hit
number
n15
from
the
enemy
library
which
basically
seemingly
targeted
curry,
and
this
turns
out
to
be
a
little
unusual
in.
But
if
you
look
at
the
saturation
of
the
pythagorean
emission
versus
n15
concentration
with
this
particular
molecule,
you
don't
get
you
get.
F
F
F
In
that
particular
case,
the
enzyme-inhibited
complex
actually
has
some
residual
activity,
so
you
can
never
push
the
system
to
complete
inhibition
and
one
possible
interpretation
of
that
is
that
the
molecule
in
question
n15,
overlaps
or
is
adjacent
to
the
active
sites
and
impedes,
but
does
not
prevent
the
access.
The
substrates
to
the
outside.
A
The
the
two
students
you
mentioned
if
they
are
thinking
about
anything
to
do
with
compound
design,
it
would
be
good
to
involve
them
in
the
in
the
project,
for
example,
on
on
github,
because
ed-
and
I
have
been
talking
a
little
bit
about
this
and
I'm
sure
joe-
is
really
interested
in
in
thinking
about
compound
design.
Building
on
the
molecules
that
have
been
obtained
so
far
just
to
try
and
loop
them
into
the
project,
that'll
be
they'll,
be
great
to
make
sure
we're
not
you
know,
duplicating
each
other.
A
C
A
G
G
I
had
a
couple
of
weeks
ago
discovered
a
new
crystal
form
with
one
molecule
of
asymmetric
unit,
which
was
much
nicer
than
the
eight
molecules
based
metric
unit
of
the
previous.
Crystal
form
goal
was
to
regrow
these
crystals
and
use
them
for
soaps.
For
the
az
compounds,
the
provider
of
our
optimization
screens,
they
had
issues
getting
a
compound
delivered
and
then
they
had
a
corona
case
in
the
office.
So
I
hope
to
get
this
green
this
week
and
then
we'll
grow
crystals
and
do
the
soaks.
C
C
A
A
B
A
A
Yes,
yes,
that's
right,
so
so
the
the
compounds
that
were
delivered-
and
you
know
what
I'm
just
gonna-
I
can't
find
the
oh
yeah.
There
you
go
yeah
these
compounds
were
delivered.
These
are
the
the
fragments
that
have
been
elaborated
further
with
some
semi-random
changes
based
on
some
of
the
previous
elaborated
fragments
that
we
got.
That
appeared
to
be
suggest.
That
appears
to
be
inhibiting
more
than
one
more
so
a
separate
line
of
inquiry,
but
the
compounds
are
looking
around.
C
A
Okay,
great
so
yes,
just
just
to
summarize,
we've
been
working
since
about
march,
I
suppose,
on
securing
molecules
predicted
to
bind
where
the
original
fragments
were
binding,
the
original
fragments
that
we
pursued
and
which
those
molecules
are
derivatives
of,
and
we
we
started
this
ai
machine
learning,
competition,
asking
people
to
use
generative
methods
to
design
new
molecules
that
might
bind
those
errors.
Someone
has
been
a
lot
to
the
previous
cause.
A
Jan
jensen
suggested
some,
and
there
were
several
other
entries
that
came
in
and
the
some
of
those
molecules
have
been
bought
and
some
of
them.
I
think
those
are
being
shipped
directly
to
laura,
where
we
can
do
that
and
others
are
being
made
by
by
ed
and
by
daniel
who's.
Also
on
the
call-
and
the
idea
is
to
complete
this
set
and
and
send
those
for
evaluation
by
laura
to
see
if
we
get
any
hits.
So
it's
competition
we're
going
to
see
what
works
best
and
everyone
here
has
used
different
methods.
A
So
it's
a
kind
of
modelling
competition
with
a
with
a
crystal
structure
as
the
as
the
target
and
and
no
other
clues.
Besides,
where
those
fragments
are
interacting.
With
the
protein
yeah
we'll
see
what
we'll
get
we
may
get,
nothing,
hopefully
we'll
get
something.
These
are
all
probably
going
to
be
weak
binders,
so
certainly
spr
and
soaking
will
be
the
best
way
to
go.
A
But
we'll
we'll
see
what
we
get.
I
guess
just
to
flag
up.
If
some
of
those
molecules
are
being
delivered
directly
to
you
from
the
commercial
suppliers
from
from
enamine,
then
I
mean
I
know
you
will
do
this,
but
but
please
keep
those
to
one
side.
We,
we
kind
of
have
a
plan
because
we're
generating
lots
of
molecules
at
the
moment
we
kind
of
have
a
plan
to
try
and
develop
a
little
bit
of
a
a
kind
of
in-house
screening
library
that
could
be
used
for
other
campaigns.
A
A
B
A
F
A
A
H
So
the
the
synthetic
process
was
extremely
weird
because
the
yeah,
you
know
you
guys-
can
see
the
pork
protected.
The
book
protected
intermediate
was
was
really
really
hard
to
separate
it.
It
got
stuck
in
the
reverse
phase
column.
It
also
cannot
come
up
from
the
normal
phase.
The
only
way
to
do
is
to
use
the
is
to
use
amino
normal
phase
column,
then
further
purify
by
hp,
further
purified
by
hp,
yeah
lcms,
basically
it's
hplc
and
then
still
cannot.
H
Sepa
cannot
separate
the
block
protected
product
with
the
starting
material,
so
I
have
to
carry
it
as
crude
to
the
next
step.
Tfa
d.
Protecting
then,
because
the
the
drastic
changing
the
polarity
of
the
compound,
then
eventually
I
got.
I
I've
been
able
to
separate
this
product
from
from
the
impurity
starting
material,
the
chloride,
the
chloride
on
the
left
hand
side
so
that
that
six
milligram
can
be
improved.
H
A
So,
just
on
this
on
this
compound,
obviously,
yes,
the
thing
we
need
to
do
is
to
check
that
it
is
active
again
in
the
enzymatic
assay
right,
and
this
would
be
this.
We
must,
because
this
is
the
analog
of
the
original
aza
compound,
where
this
nh2
is
an
oh.
So
I
guess
we're
we're
keen
to
try
to
send
some
to
work
to
make
sure
it
works,
and
then,
if
it
does
have
inhibitory
activity,
then
the
next
thing
would
be
to
test
it
on
on
bacteria,
yeah,.
H
D
C
D
A
Relationship,
yes,
yeah.
These
are
structural,
elicitation
things,
so
we're
trying
to
get
structures
of
these
compounds
by
micro,
ed,
that's
all
to
try
and
accelerate
how
we're
doing
structural
elucidation,
but
it's
not
directly.
It's
not
directly
relevant
to
the
stuff
we're
doing
in
finding
bioactive
compounds.
It's
just
that
we're
trying
to
accelerate
things.
A
C
A
H
H
G
H
A
A
A
A
D
We've
had
all
of
the
structures
sent
to
us
and
the
supporting
information
will
be
on
the
way
as
well.
So
that's
amazing,
that's
happening
so
we'll
update
next
time
once
I've
actually,
once
we've
actually
got
them
all,
to
give
you
a
list
of
what's
arrived
and
what
they
look
like.
So
we'll
assemble
we'll
assemble
that
into
something.
So
people
can
have
a
look
at
what
the
structures
are
and
what's
to
come
through
from
life
arc,
but
they
they
will,
they
will
feed
into
the
assay
pipeline.
D
The
second
thing
about
the
assay
pipeline
is
that
we
were
held
up
with
the
stopped
with
the
continuous
essay
that
we
have
for
the
data
analysis.
So
we've
got
an
it
project
on
at
the
moment
this
week
to
get
a
nice
little
script,
to
be
able
to
get
the
kinetic
constants
out
of
the
the
continuous
assay
to
be
able
to
identify
where
the
linear
slope
is
etc
and
work
back
from
that
the
inhibition.
D
So
that's
ongoing
this
week,
so
we're
hoping
that
our
throughput
will
accelerate
quite
substantially
soon
with
the
next
week
or
so
in
in
advance
of
any
advances
on
the
stopped
essay.
I
mean
to
be
honest:
we
need
another
pair
of
hands.
I
mean
that's,
I'm
writing
furiously
for
fellowships
at
the
moment
and
to
get
more
peasant
hands
all
sorts
of
places.
So,
but
anyway
it's
it's.
A
A
Last
time,
we'd
we'd
agreed
to
wait
to
reach
out
to
cc4
carb
until
we'd
perhaps
got
some
fleshed
out
some
of
the
more
recent
data
so,
for
example,
getting
a
structure
and
so
on,
and
I
think
chris
you
were
chasing
on
the
global
global
health
library
in
case
that
comes
through.
I
think
that
was
a
contract.
A
There
is
something
which
isn't
up
here,
which
is
on
a
parallel
project
that
we
have
going
on
a
fungal
infection
that
unesco
are
asking
for
written
submissions
on
open
source
projects
as
exemplars
of
how
to
work
with
open
science.
The
deadline
is
like
friday.
It's
a
pretty
simple
form.
I
need
to
submit
one
for
a
couple
of
other
projects,
I'll
try
and
submit
one
for
this
project
and
I'll
try
and
post
the
text
somewhere
so
that
everyone
can
see
it.
A
A
Yeah,
I
think
you're-
probably
right,
but
yeah
you're,
probably
right,
but
I
I
guess
I'm
keen
to
submit
something,
because
there
are
quite
a
lot
of
open
science
projects
which
are
ultimately
about
open
access
and
things
like
metadata
and
stuff
like
that,
and
there
are
actually
relatively
few
open
science
projects
which
are
actually
doing
it
in
the
way
that
we're
doing
it.
So.
C
A
A
I'm
also
so
in
a
couple
of
weeks,
I'm
giving
a
talk
at
the
gordon
conference
on
new
antibacterial
agents.
It's
a
week-long
thing
in
italy
easily
I'm
on
the
last
day,
so
I'll
be
able
to
listen
everyone
else's
talks
and
then
and
then
talk
about
what
we're
doing
again,
I'll
really
try
and
share
a
deck
before
that
or
share
it
around.
So
people
are
happy
with
it.
A
The
aim,
of
course,
is
to
acknowledge
everyone
in
full,
so
I'm
I'm
speaking
only
on
behalf
of
this
group
and
also
on
behalf
of
another
group
series,
two
in
open
source
antibiotics
which
is
not
related
to
this
one.
But
again
I'm
not
saying
anything
which
is
not
already
public,
and
I
will
be
very,
very
careful
to
make
sure
that
everyone's
acknowledged
suitably,
but
that
that
talk
is
what
is
it
the
20
28
or
something.
C
A
D
A
A
Interestingly,
the
the
that
meeting
was
cancelled
right
at
the
beginning
of
the
pandemic
in
2020,
because
there
was
a
spike
in
cases
around
naples,
which
is
where
the
conference
was
so
it
was
the
first
thing
I
saw
of
to
be
cancelled
and
it's
due
to
start
in
a
couple
of
weeks
and
I'm
just
hoping
the
case
is
stacked
down.
To
avoid
a
grotesque
repetition
of
history.
